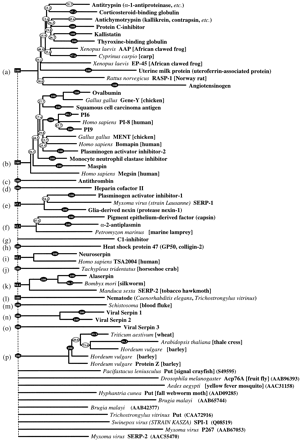
Multifurcating phylogenetic tree indicating the overall relationship between members of the serpin superfamily. The tree is a combination of the majority consensus maximum parsimony trees seen in Figure 4, with groups of serpins of similar type (e.g., antithrombin) represented by a single identifier, where possible. The branch lengths reflect maximum likelihood distances introduced using the method of Fitch and Margoliash (1967), as implemented in FITCH (Felsenstein 1996). Conventional bootstrap values from the maximum parsimony trees appear as ovals, rectangles indicate those subtrees whose members were identified using the comparison method, and hexagons indicate those identified by the strict consensus method. The 10 orphans are at the bottom of the tree. Clade identifiers (a, b,c, etc.) are in parentheses and correspond with subgroups identified in Figure 4, Table 3, and the text.











