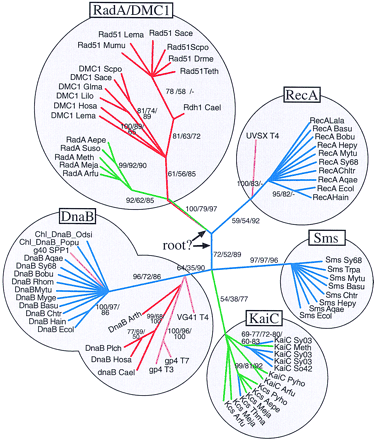
Unrooted phylogeny of the RecA/DnaB superfamily. The analysis is based on an the alignment of the RecA/DnaB core domain shown in Fig. 1. The data matrix contains 221 residues seven of which are invariant or parsimony uninformative. Support for individual branches is indicated by bootstrap values for 1000 resampling of PAUP maximum parsimony (first number), PHYLIP distance analysis (second number), and the reliability value computed by the PUZZLE software (third number). Bootstrap values <50% are not recorded and branches without bootstrap numbers are derived from a distance tree computed with the PHYLIP programs protdist and fitch. Branch lengths are arbitrary and do not represent evolutionary distances. The two possible positions of the root as discussed in the text are indicated by black arrows. (Red) Eukaryota; (green) Archaea; (blue) Bacteria; (pink) Bacteriophages. Names in boxes identify the individual protein families. The sequence identifiers are the same as for Fig. 1 except that the GenBank identifier was omitted.











