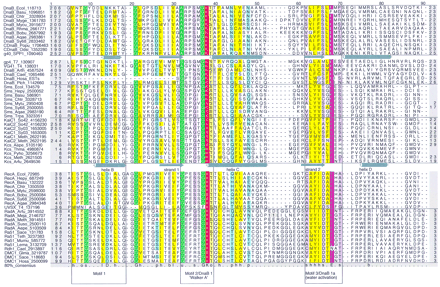
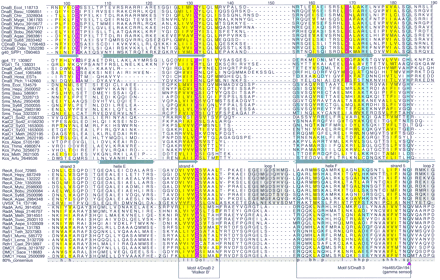
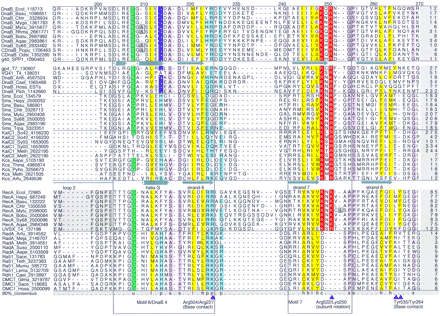
Multiple alignment of the core domain of the RecA/DnaB superfamily of ATPases. From top to bottom (separated by horizontal lines) the alignment contains sequences from bacterial and chloroplast DnaB, DnaB proteins and primase–helicase proteins from bacteriophages and eukaryotes, bacterial Sms proteins, KaiC from Archaea and Bacteria, RecA recombinase from Bacteria and phage T4, and RadA and Rad51/DMC1 recombinases from Archaea and Eukaryota. The 80% consensus for these proteins is shown below the aligned sequences. Numbers indicate the distance to the amino-terminal methionine and the carboxyl terminus of each protein and residues omitted within the alignment. (&) The position of inteins that have not been included in the alignment. The secondary structure elements derived from the X-ray structures of phage T7 gp4 and E. coli RecA are shown above the respective sequence. Helices are represented as cylinders, strands as arrows, and the unordered or mobile loops 1 and 2 as lines. Key residues that are discussed in the text are marked by arrowheads; the numbers identify the position of the residue in gp4 and RecA according to the original publications (Story et al. 1993; Sawaya et al. 1999). Highly conserved residues are color coded and indicated in the consensus line for the following groups. (Purple) Negatively charged (D,E); (red) positively charged (H,K,R), charged (c = D,E,H,K,R); (green) tiny (u = G,A,S); (yellow) hydrophobic (h = A,C,F,I,L,M,V,W,Y) or aliphatic (l = I,L,V); (pale yellow) alcohol (o = S, T, Y); (light blue) polar (p = D,E,H,K,N,Q,R,S,T), (reddish-brown) small (s = A,C,D,G,N,P,S,T,V); (gray) big (b = not small). Also colored are residues conserved only within the DnaB family. Where applicable, source organisms are identified by four-letter abbreviations. (Aepe) Aeropyrum pernix; (Aqae)A. aeolicus; (Arfu) Archaeoglobus fulgidus; (Arth)A. thaliana; (Basu) Bacillus subtilis; (Bobu)Borrelia burgdorferi; (T7) bacteriophage T7; (T4) bacteriophage T4; (Cael) C. elegans; (CDnaB_Odsi)Odontella sinensis chloroplast; (CDnaB_Popu) Porphyra purpurea chloroplast; (Chtr) Chlamydia trachomatis; (Ecol)E. coli; (Glma) Glycine max; (Hain) Haemophilus influenzae; (Hepy) H. pylori; (Hosa) Homo sapiens; (Lema) Leishmania major; (Meja)Methanococcus jannaschii; (Meth) Methanobacterium thermoautotrophicum; (Mumu) Mus musculus; (Myge)Mycoplasma genitalium; (Mytu)Mycobacterium tuberculosis; (Plch) P. chabaudi; (Rhma) Rhodothermus marinus; (Sace) Saccharomyces cerevisiae; (SPP1)Bacillus subtilis bacteriophage SPP1; (Suso) Sulfolobus solfataricus; (Sy68) Synechocystis PCC6803; (Teth)Tetrahymena thermophila; (Thma) T. maritima; (Trpa)Treponema pallidum.











