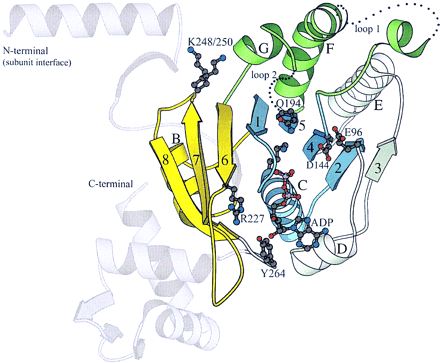
A molscript diagram of E. coli RecA structure. Areas with sequence conservation between DnaB and RecA/DMC1/RadA are highlighted, the nonconserved carboxy- and amino-terminal domain are shown in light gray. The central parallel μ-sheet is blue and the elements that are involved in coordinating loop 1 (between strand 4 and helix F) and loop 2 (between strand 5 and helix G) are green. Areas within the core domain that show no obvious sequence similarity between RecA and DnaB (helix D, strand 3 and helix E) are shown in light blue. The subdomain composed of helix B and strands 6–8 is shown in yellow. ADP diffused into the crystal (Story and Stitz 1992) is shown in ball-and-stick representation. Conserved amino acid residues that are discussed in the text and indicated in the alignment (Fig. 1) are shown in ball-and-stick representation: Lys-72 (in the P-loop), Glu-96, Asp-144 (Walker B), Gln-194, Arg-227, Lys-248, Lys-250, and Tyr-264 are at or near the carboxyl terminus of strands 2, 4, 5, 6, and 7, respectively. Amino acid coordinates are from PDB file 2REB, location of ADP is from PDB file 1REA. The orientation of the monomer, labels of strands, helices, loops, and residue enumeration are in accordance with the original publications (Story et al. 1992; Story et al. 1993).











