Overview of MCL clusters
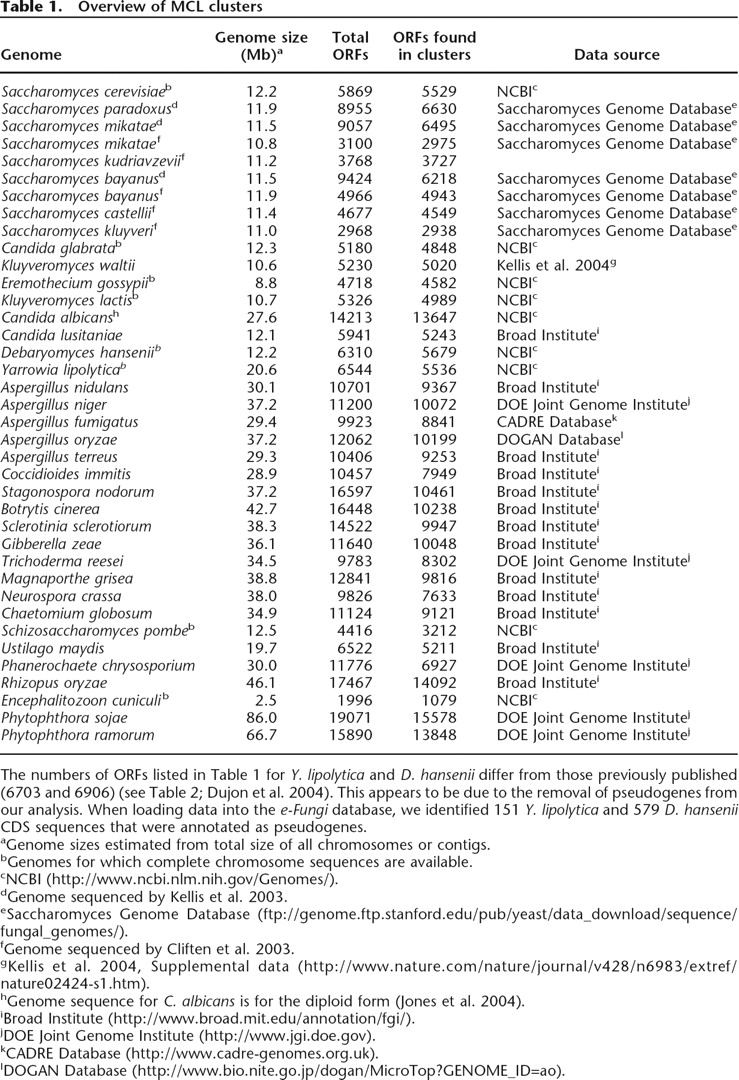
The numbers of ORFs listed in Table 1 for Y. lipolytica and D. hansenii differ from those previously published (6703 and 6906) (see Table 2; Dujon et al. 2004). This appears to be due to the removal of pseudogenes from our analysis. When loading data into the e-Fungi database, we identified 151 Y. lipolytica and 579 D. hansenii CDS sequences that were annotated as pseudogenes.
aGenome sizes estimated from total size of all chromosomes or contigs.
bGenomes for which complete chromosome sequences are available.
cNCBI (http://www.ncbi.nlm.nih.gov/Genomes/).
dGenome sequenced by Kellis et al. 2003.
eSaccharomyces Genome Database (ftp://genome.ftp.stanford.edu/pub/yeast/data_download/sequence/fungal_genomes/).
fGenome sequenced by Cliften et al. 2003.
gKellis et al. 2004, Supplemental data (http://www.nature.com/nature/journal/v428/n6983/extref/nature02424-s1.htm).
hGenome sequence for C. albicans is for the diploid form (Jones et al. 2004).
iBroad Institute (http://www.broad.mit.edu/annotation/fgi/).
jDOE Joint Genome Institute (http://www.jgi.doe.gov).
kCADRE Database (http://www.cadre-genomes.org.uk).
lDOGAN Database (http://www.bio.nite.go.jp/dogan/MicroTop?GENOME_ID=ao).