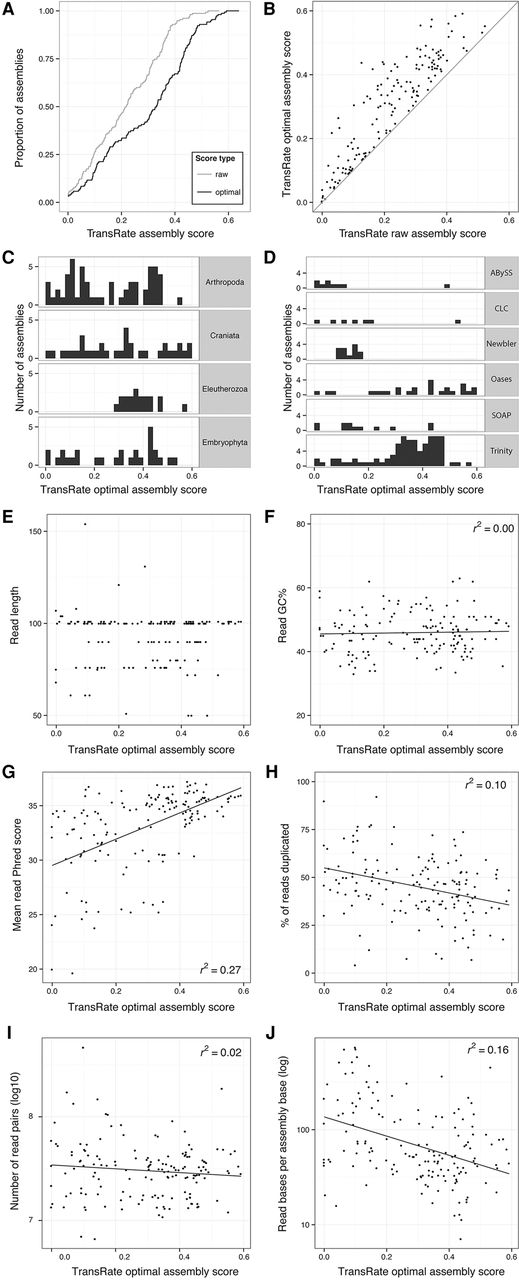
Application of TransRate to 155 published assemblies from the NCBI Transcriptome Shotgun Archive. One hundred fifty-five assemblies from the Transcriptome Shotgun Archive were analyzed using TransRate. The quality of the reads used to generate the assemblies were also analyzed using FastQC. (A) Cumulative distribution of TransRate raw and optimized assembly scores for each of the 155 assemblies. (B) Comparison between raw and optimized assembly score. (C) Distribution of TransRate optimized assembly scores partitioned by taxonomic group. (D) Distribution of TransRate optimized assembly scores partitioned by assembly method. (E–J) TransRate optimized assembly scores compared to various summary statistics of the input reads: (E) read length; (F) read GC%; (G) mean read per-base Phred score; (H) percent of reads that were PCR duplicates; (I) number of read pairs; and (J) read bases per assembled base.