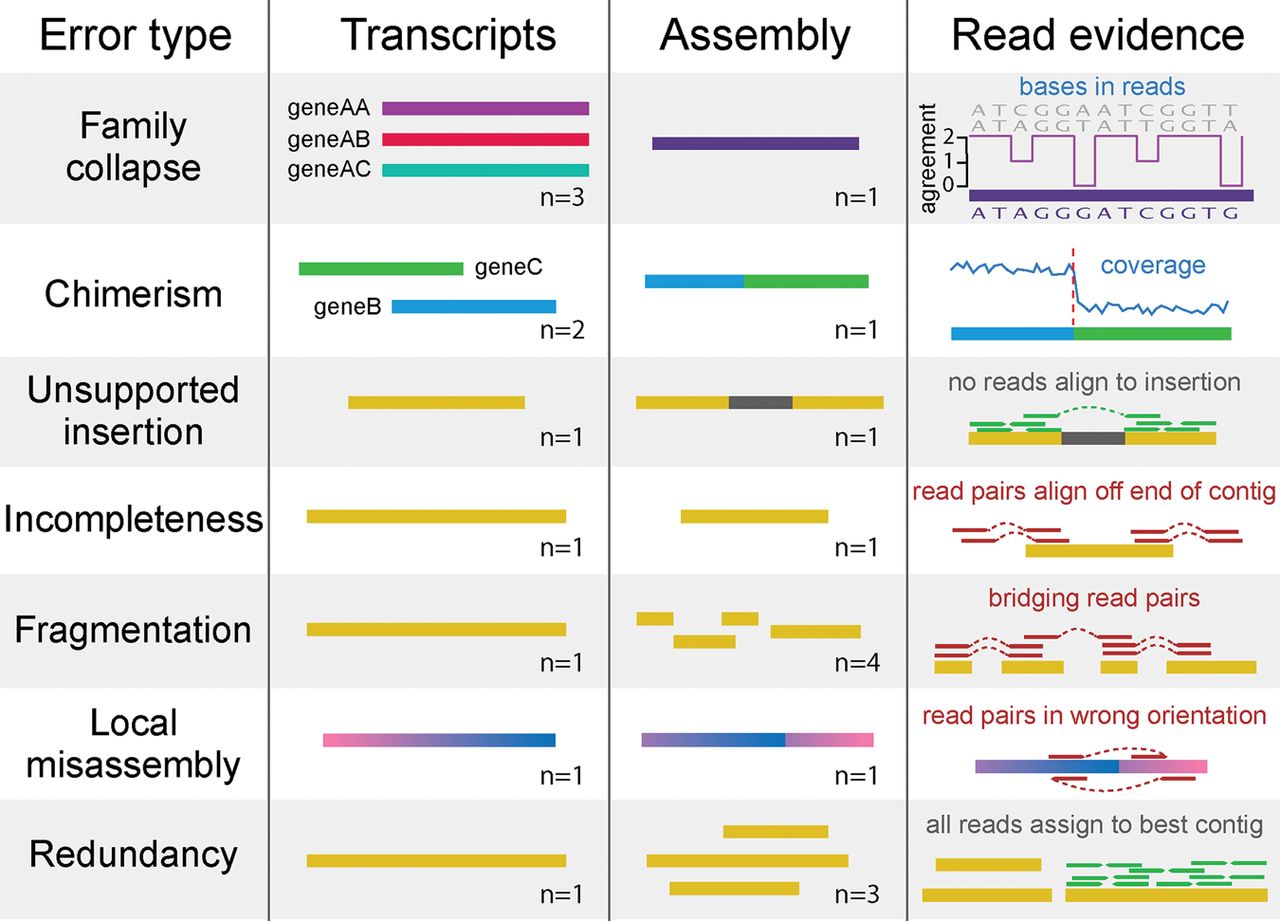
Common errors in de novo transcriptome assembly, and how they can be detected using read mapping data. Family collapse occurs when multiple members of a gene family are assembled into a single hybrid contig. This error can be detected by measuring the extent that the nucleotides in the contig are supported by the mapped reads. Chimerism occurs when two or more transcripts (that may or may not be related) are concatenated together in a single contig during assembly. This can be detected when the expression levels of the transcripts differ, leading to a change-point in the read coverage along the contig. Unsupported insertions can be detected as bases in a contig that are unsupported by the read evidence. Incompleteness can be detected when reads or fragments align off the end of the contig. Fragmentation is caused by low coverage and is detectable when read pairs bridge two contigs. Local misassembly encompasses various structural errors that can occur during assembly, such as inversions, usually as a result of assembler heuristics. These are detectable when both members of a read pair align to a single contig, but in a manner inconsistent with the sequencing protocol. Redundancy occurs when a single transcript is represented by multiple overlapping contigs in an assembly. This is detectable when reads align to multiple contigs but the assignment process assigns them all to the contig that best represents the original transcript.