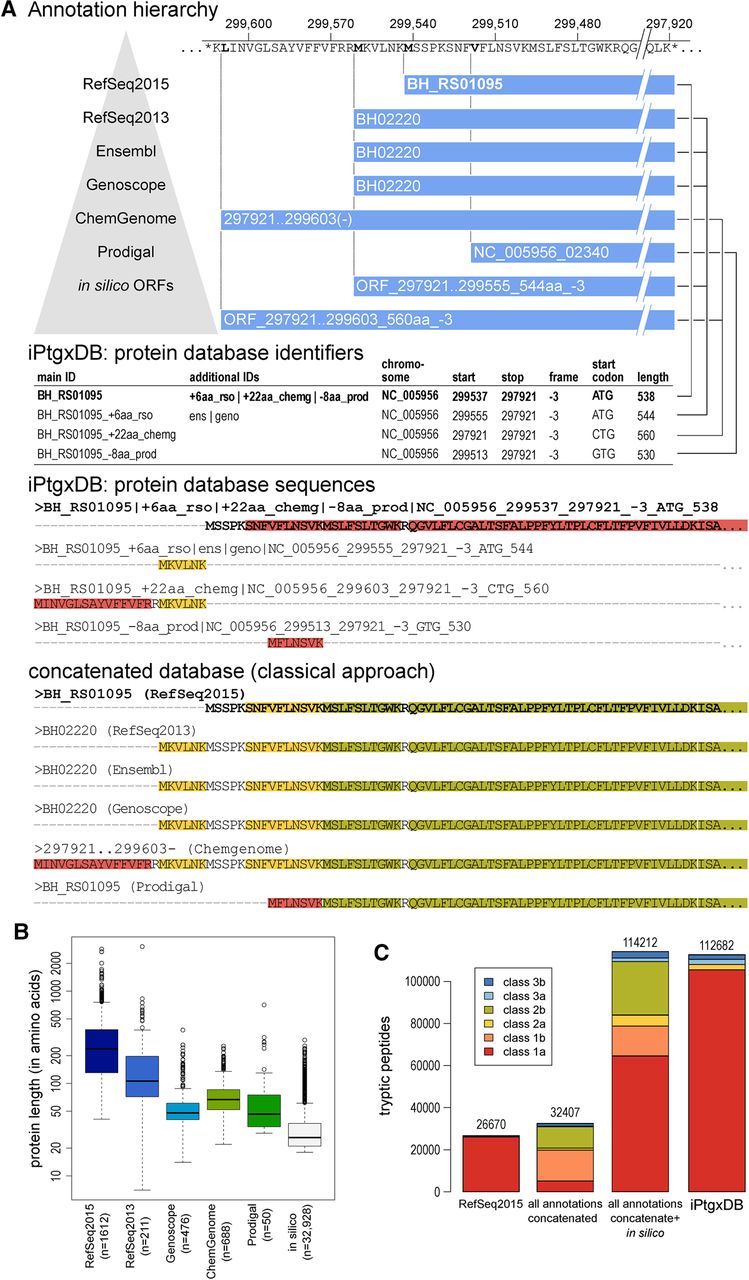
Generating an iPtgxDB with informative identifiers and a minimally redundant protein search DB in FASTA format. (A) CDSs and pseudogenes of seven resources are integrated in a stepwise fashion. Informative protein identifiers are created and illustrated for the annotation cluster, with the RefSeq2015 anchor sequence BH_RS01095 shown in bold, where three additional start sites exist. The four different proteoforms are added to the protein search DB: the anchor sequence (bold) with the full protein sequence, the extensions (RefSeq2013 and ChemGenome) add the upstream sequence up to the first tryptic cleavage site within the anchor sequence. The shorter Prodigal prediction uses an alternative start codon resulting in a distinguishable N-terminal peptide and therefore also gets added. The two in silico ORFs are identical to annotations higher up in the annotation hierarchy and therefore are not added. Peptide classes are shown for the N-terminal sequences of the CDS annotation cluster (see also Fig. 2C). (B) Box plots of protein length for RefSeq2015 and those proteins that get added in each successive step to the protein search DB illustrate that we include many sORFs potentially missed in the reference annotations. (C) Bar chart showing the DB complexity and the peptide classes for RefSeq2015, all six integrated annotations without and with in silico ORFs, and the final iPtgxDB. The legend (inset) shows colors for the six peptide classes.