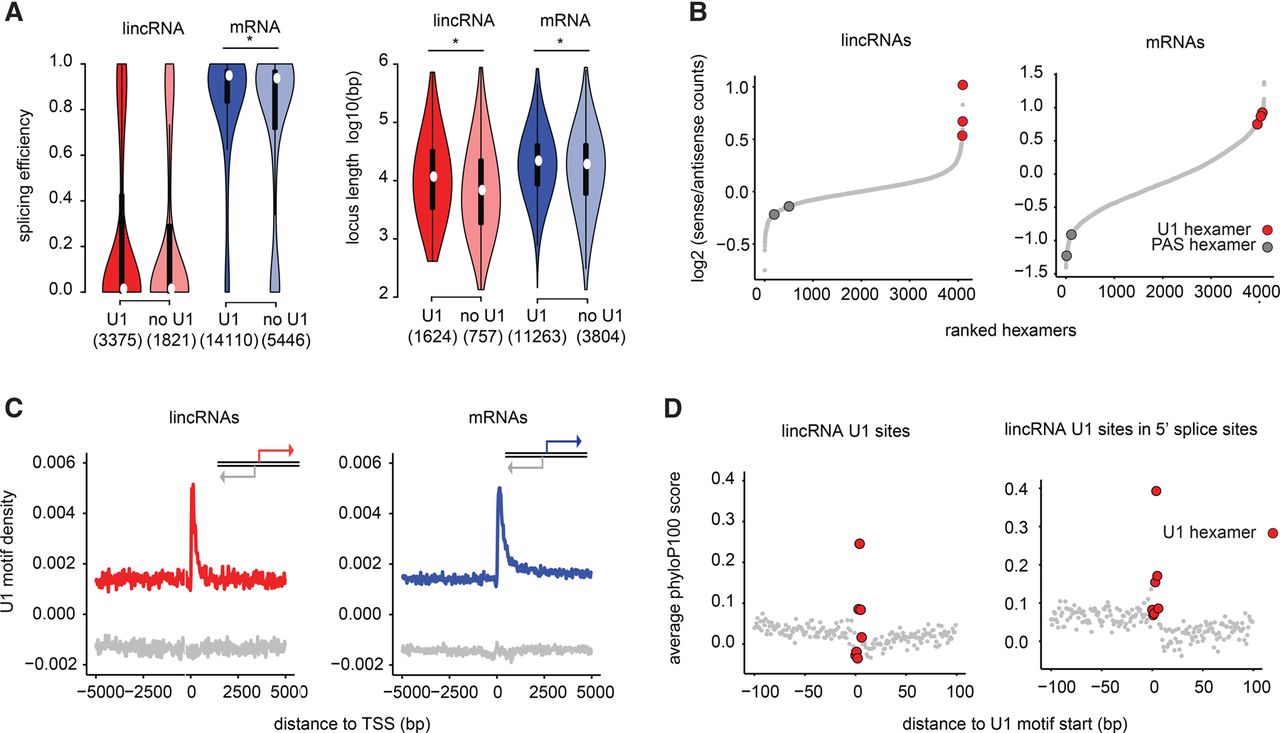
The U1-PAS axis is similar in lincRNAs and mRNAs. (A) Distribution of splicing efficiency (left) and locus length (right) in lincRNAs and mRNAs in K562, with and without canonical U1 motifs at 5′ splice sites within the first 1 kb downstream from the TSS. Asterisks indicate significant differences (Wilcoxon P-value <0.05). (B) Rank of all hexamers by enrichment in the first 1 kb in the sense direction relative to upstream antisense direction in lincRNAs (left) and mRNAs (right). (C) U1 motif density around the TSS in lincRNAs (left) and mRNAs (right) in the sense strand (red or blue) and in the antisense strand (gray). (D) Average conservation at all U1 nucleotides present in the first 1 kb of lincRNAs (left) or the subset of these U1 nucleotides that overlap 5′ splice donors (right).