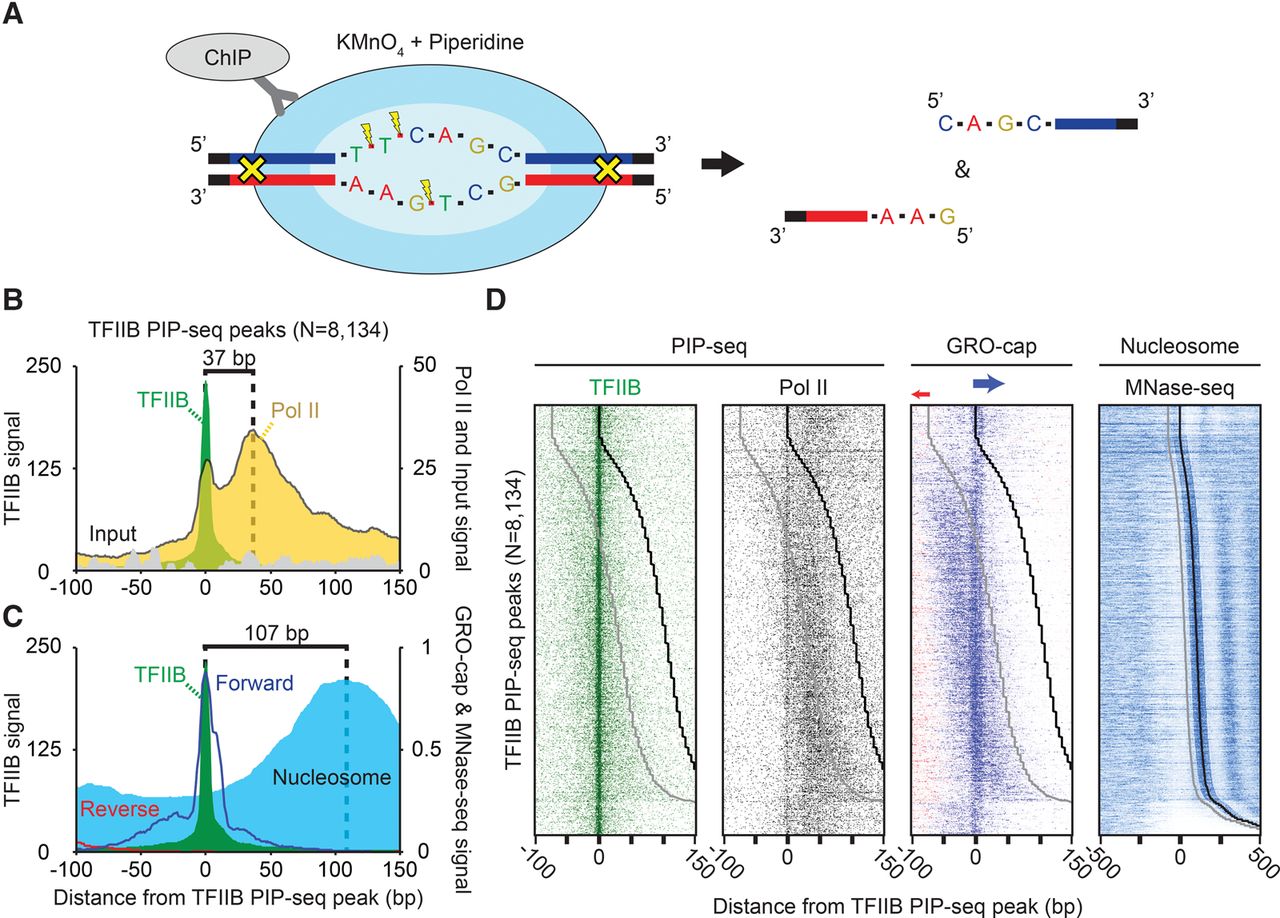
Positional separation of open preinitiation complexes (PICs) and paused complexes (PCs) associated with ncRNA and mRNA transcription. (A) Schematic of the PIP-seq assay. KMnO4 oxidizes single-stranded thymines, which are subsequently cleaved by piperidine. Coupled to formaldehyde-based crosslinking and immunoprecipitation, open DNA relative to a protein of interest is enriched. (B) Composite plots of PICs (N = 8134). TFIIB-bound open complexes were identified as enriched TFIIB PIP-seq peaks (see Methods) that also had a corresponding enrichment of TFIIB ChIP-exo peaks, as well as GRO-cap transcription. (C) Composite plots of PICs (N = 8134) overlaid with composite plots of GRO-cap (RNA) and MNase-seq tag 5′ ends (nucleosomes) that were shifted in the 3′ direction by 80 bp or approximately half the average fragment length (Lai et al. 2012; The ENCODE Project Consortium 2012; Core et al. 2014). (D) Heatmap of PICs (N = 8134), sorted by the distance between TFIIB PIP-seq peaks to downstream +1 nucleosomes. The black line represents the consensus +1 nucleosome dyad, and the gray line is 73 bp upstream of the dyad representing the upper edge of the nucleosome. The few dyads that align exactly on the TSS are likely artifacts resulting from parameter settings and thresholding and thus should be ignored.