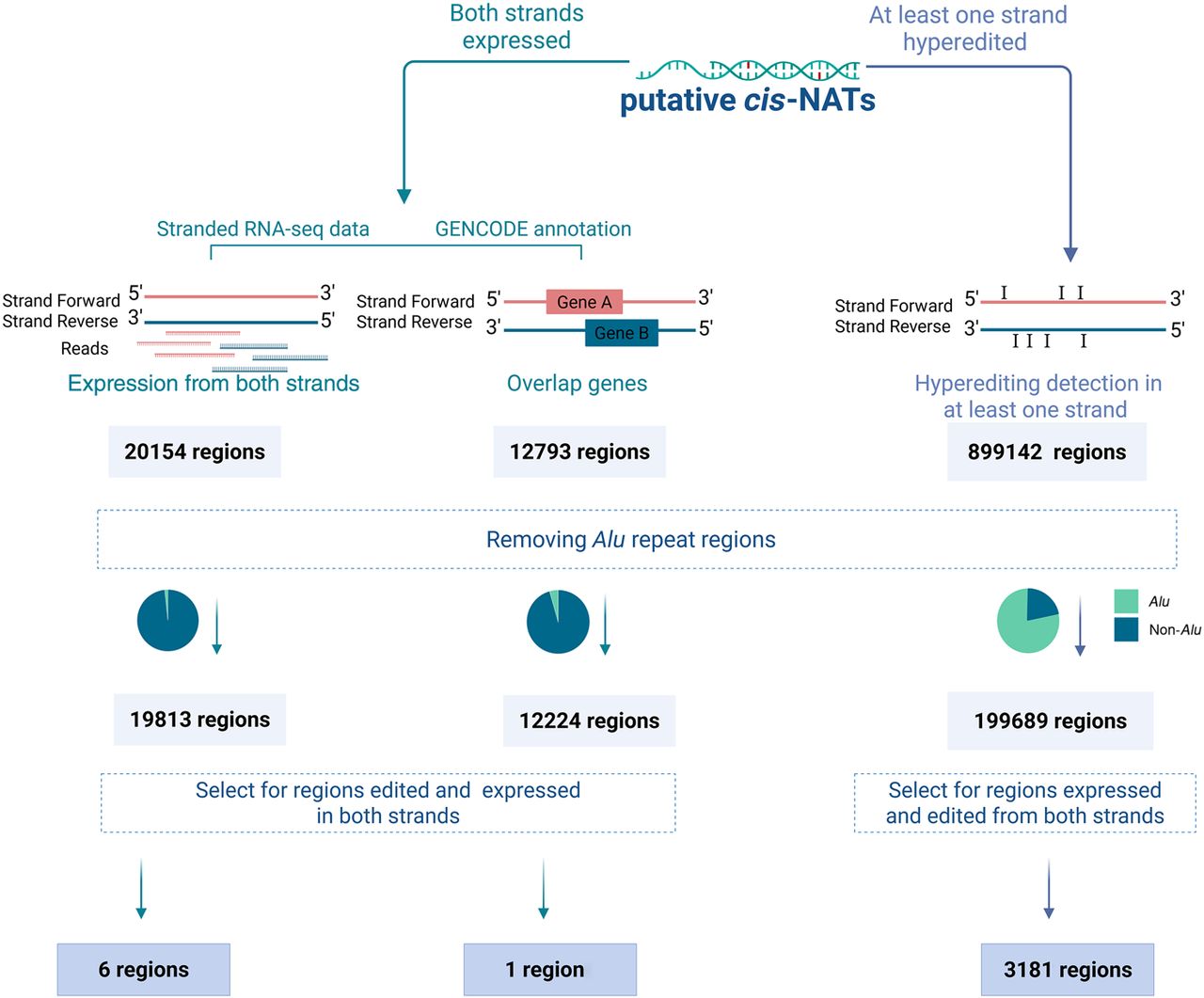
Figure 2.
Defining putative cis-NAT regions using three approaches. Putative cis-NATs were identified in two ways: (1) searching for genomic regions expressed from both strands using stranded RNA-seq expression data (left) and gene annotation data (middle) and (2) searching GTEx data for regions that are hyperedited on at least one strand (right). Alu regions were excluded as they are likely to form dsRNA structures due to two oppositely oriented elements on the same strand. Regions exhibiting expression of both strands and editing levels exceeding the background were considered putative cis-NAT regions.