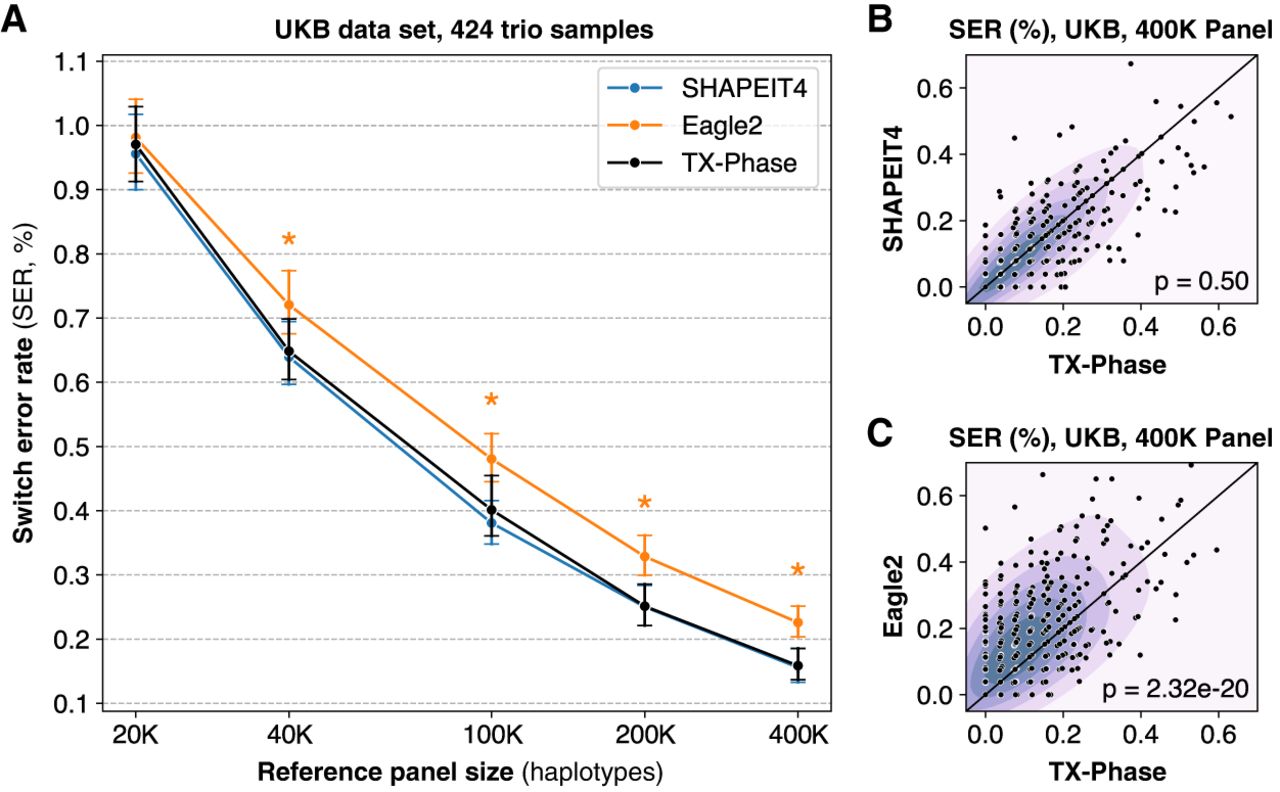
TX-Phase obtains state-of-the-art phasing accuracy on the UK Biobank (UKB) data set. (A) We compared the accuracy of TX-Phase with two standard phasing methods, SHAPEIT4 (Delaneau et al. 2019) and Eagle2 (Loh et al. 2016), on 424 trio samples from the UKB (genotyping array data; Chromosome 20) using subsets of the remaining cohort of varying sizes as the reference panel. Accuracy is measured by switch error rates (SERs) with trio-based phasing as the ground truth. Markers indicate the mean SERs, and error bars represent 95% confidence intervals. Asterisks denote highly significant differences (two-sided Wilcoxon signed-rank test, P < 1 × 10−10) between TX-Phase and Eagle2. The same statistical tests were performed between TX-Phase and SHAPEIT4, which did not result in any significant difference. We also compare the per-sample SERs of TX-Phase against those of SHAPEIT4 (B) and Eagle2 (C) for the 400,000 reference panel. The background heatmap visualizes marker density. For clarity, the display region is limited to [0, 0.7] for both axes. We show the P-values from the two-sided Wilcoxon signed-rank test, which suggest that TX-Phase significantly outperforms Eagle2 while obtaining comparable accuracy with SHAPEIT4. Consistent results are observed when phasing a sample from the Genome-in-a-Bottle Consortium using the 1000 Genomes and Haplotype Reference Consortium reference panels (Supplemental Figure S1).