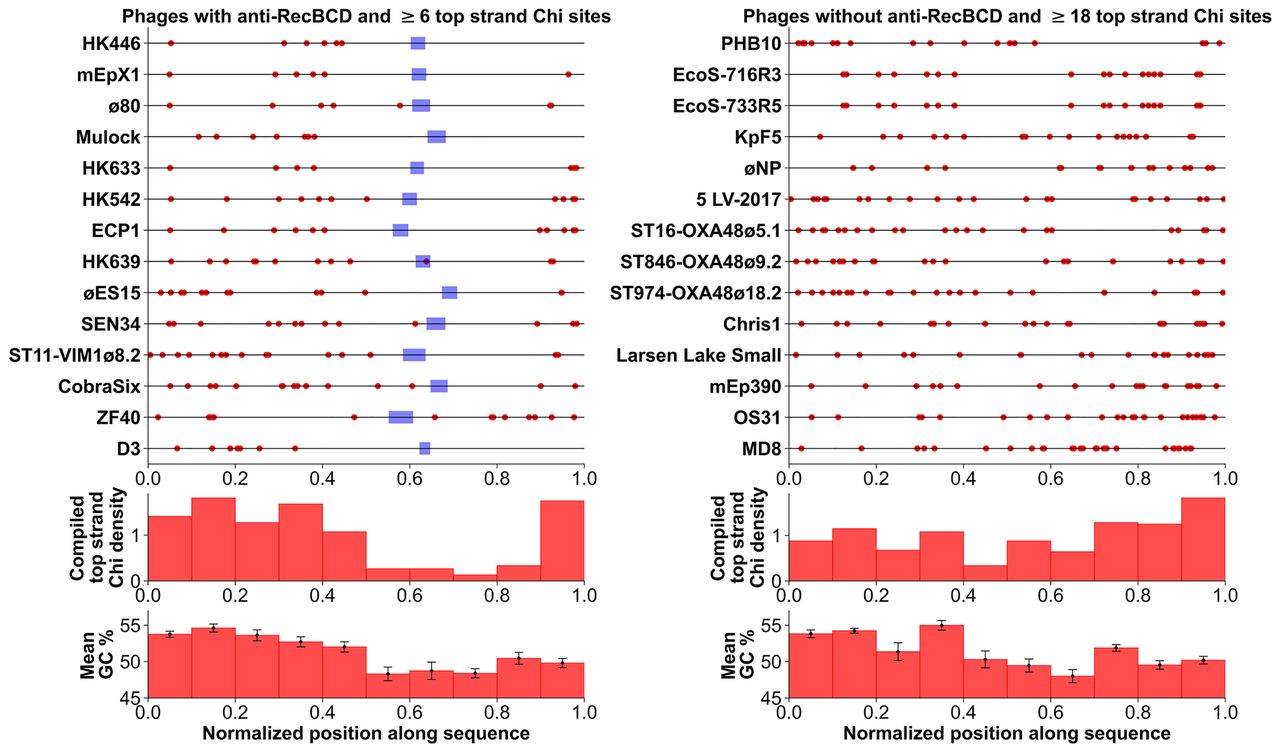
Rare, exceptional phages with both anti-RecBCD and many Chi sites often lack Chi near the Rec module. The positions of Chi sites (red dots) are placed on the genome sequence of the indicated phage as a fraction of the genome length, to facilitate comparison of genomes with different lengths. (Left) All 13 Enterobacteriales phages in the lambdoid panel (Supplemental Table S3) with an encoded anti-RecBCD function and six or more Chi sites are plotted; their Rec module is blue. (Right) All 13 Enterobacteriales phages in Supplemental Table S3 without an encoded anti-RecBCD function and 18 or more Chi sites are plotted. P. aeruginosa phages MD8 and D3 are included for comparison but are not included in the histograms of G+C content because of their high G+C content reflecting that of their host P. aeruginosa. The histogram directly below each Chi position plot shows the density of Chi sites in each 10% bin of the genome. Note that in the left panel there are many fewer Chi sites near the Rec module than far to one side or the other. The distributions in the left and right panels are significantly different (P < 0.0001 by χ2 test). This outcome is consistent with this region having recently come from a typical lambdoid phage, such as λ or P22, with a Rec module but without Chi sites (Supplemental Table S3). The histogram below each Chi density histogram shows the mean G+C content in each 10% bin of the genome, with error bars representing standard error. In both the left and right panels, the G+C content of the rightmost three bins is significantly less than that of the leftmost seven bins (P < 0.001 by paired two-sample t-test).