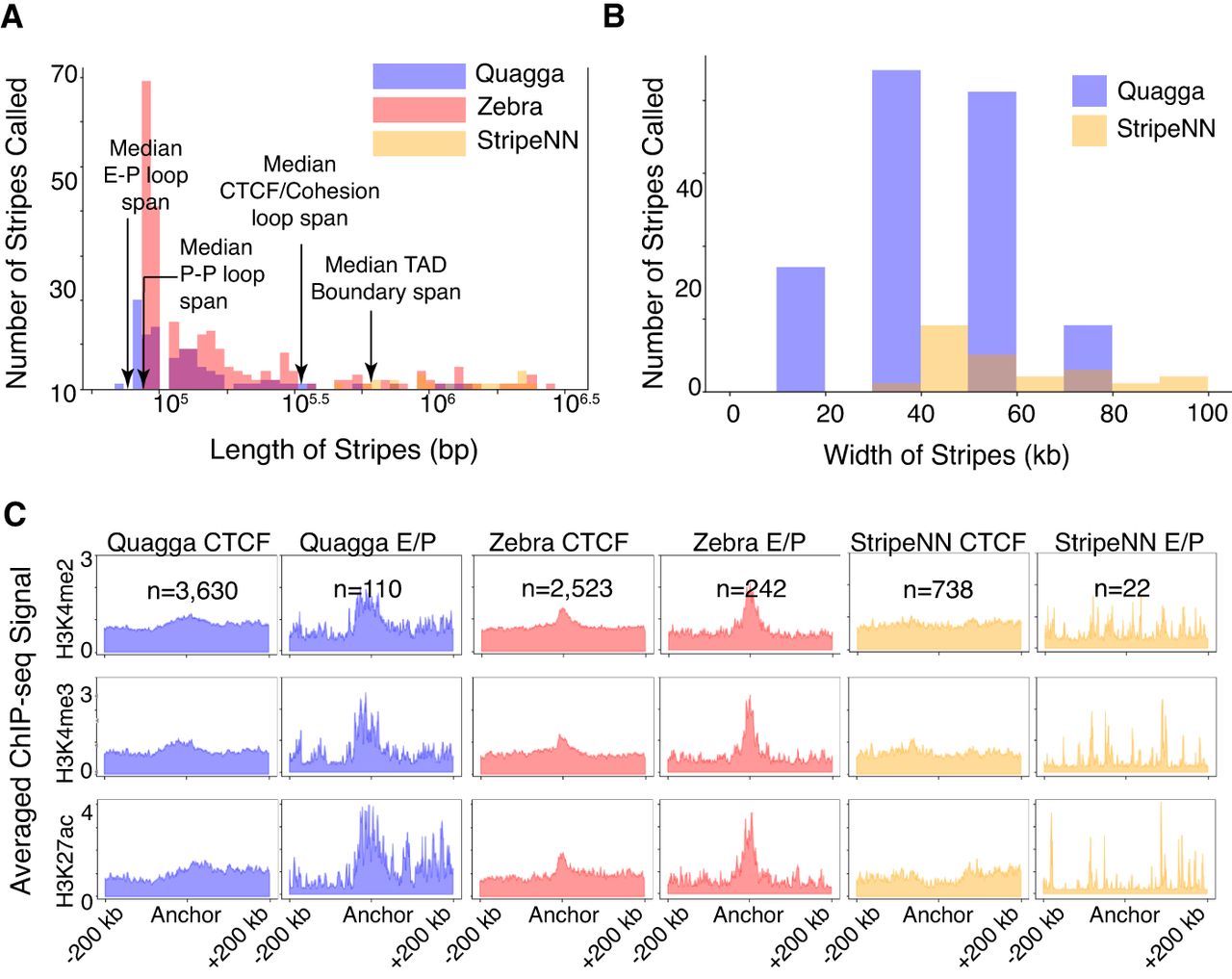
Figure 5.
Comparison of stripes called for Quagga, Stripenn, and Zebra were compared for their ability to call EP interaction–related stripes from GM12878 Hi-C data. The relevant EP-stripes were determined using ChromHMM annotations and unoccupied CTCF motifs. From filtered stripes, their distributions of stripe length (A) and stripe width (B) were determined. (C) Comparison of enrichment of EP-associated ChIP-seq signals between stripe calling tools; CTCF-occupied stripes are compared to EP-stripes on an averaged ChIP-seq pileup over a window of ±200 kb from the stripe width's coordinate mid-line for H3K4me2, H3K4me3, and H3K27ac, which are associated with EP interactions.