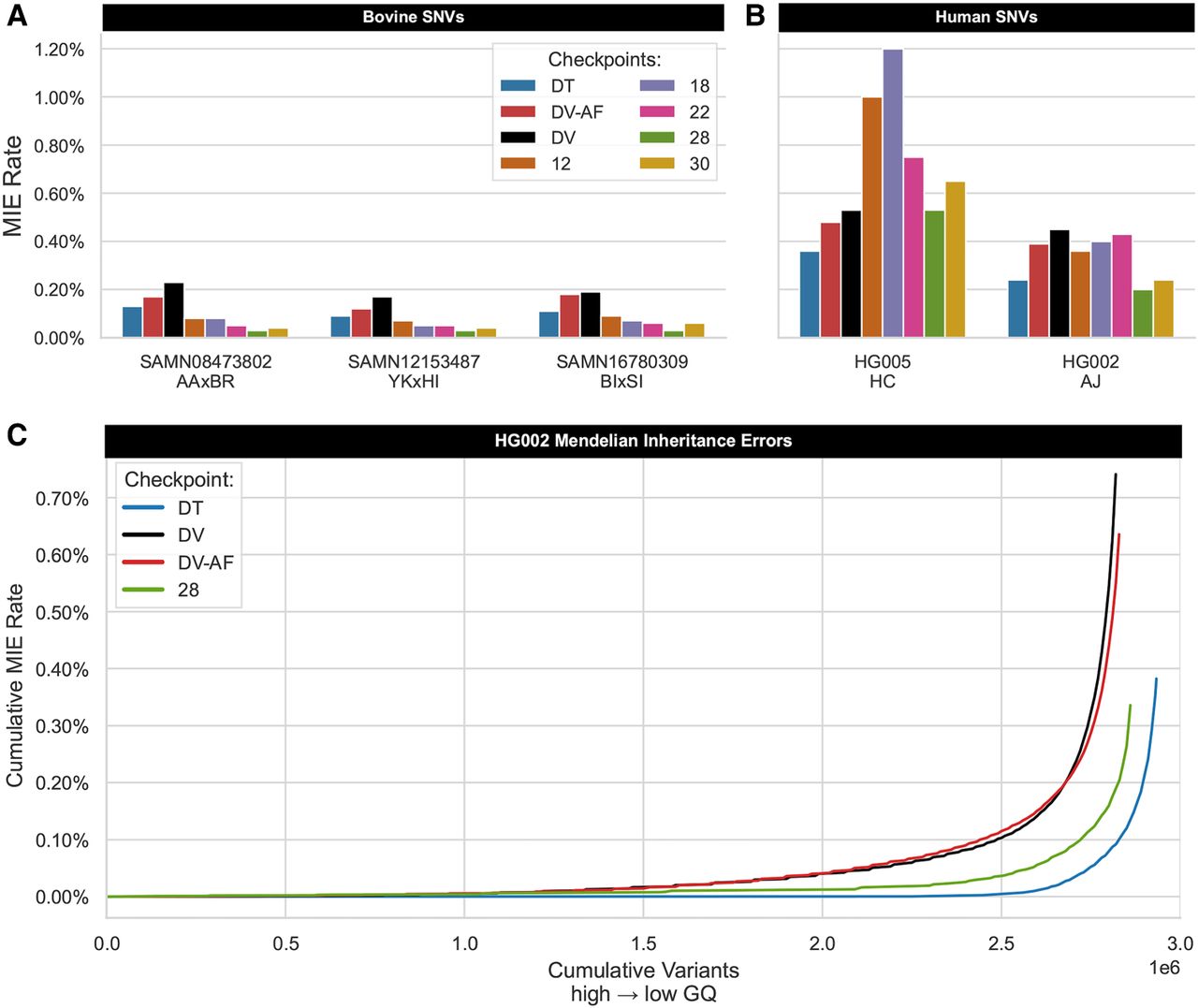
Assessing inheritance error rate in human and bovine trios. Mendelian inheritance errors (MIEs) were identified in PASS variants from the autosomes and X Chromosome for three bovine hybrid-cross trios (A) and two GIAB human trios (B). Each cluster of bars on the x-axis represents a distinct trio, with a bar representing the total number of discordant SNVs proportional to the total number of variants analyzed for eight distinct checkpoints. The Han Chinese (HC) trio deviates from all other trios, with the MIE rate nearly doubling for the first two TrioTrain phases. However, relative to the default checkpoint (DV), one bovine-AF checkpoint (28) achieves an identical SNV MIE rate in HG005 (0.53%) while also producing fewer discordant SNVs in the Ashkenazi (AJ) trio. Because checkpoints differ in total variants called in the sample individual, the line plot (C) enables a more direct comparison for HG002 only. The y-axis reports the cumulative number of errors found across four distinct versions of DeepVariant (DV, DV-AF, DT, C28). PASS variants in the trio VCF were sorted by genotype quality (GQ) in the offspring, where GQ score decreases as the number of variants increases from left to right on the x-axis. Compared with the human-trained single-sample variant callers, the error curve with checkpoint 28 approaches DT, confirming that our approach successfully recognizes inherited variation. Although DT is a joint caller requiring a minimum of two samples with known pedigree, the bovine-trained checkpoint built with TrioTrain reduces the MIE rate in individual samples, enabling speed improvements.