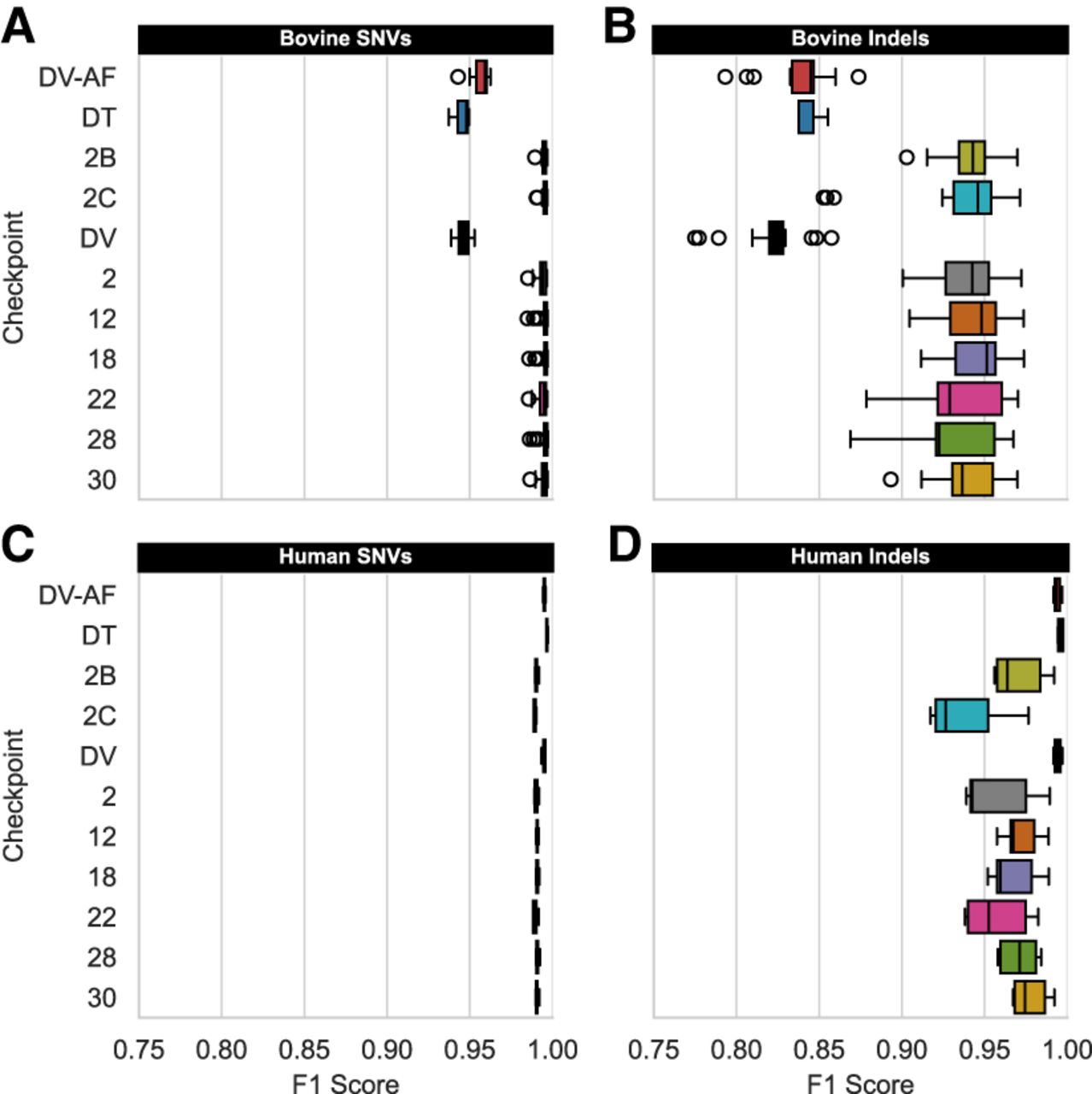
Comparing variant type generalization across species. Three human-trained versions of DeepVariant (DV, DV-AF, DT) are contrasted against eight bovine-trained DV-AF checkpoints created with TrioTrain. Each box-and-whisker represents the distribution of F1 scores observed in bovine testing genomes (A,B; N = 19, except for DT, where N = 3 offspring) or the human GIAB trios (C,D; N = 6). As expected, exclusively training with the human GIAB samples achieves a higher F1 score with these samples. Although bovine-trained checkpoints ignore genuine indels in humans, the high-quality SNV variants within the UMAGv1 callset enable SNV generalization across species. However, bovine genomes (A,B) results quantify genotyping changes relative to the GATK-based truth (UMAGv1). For results stratified by genotype class, see Supplemental Figure S15.