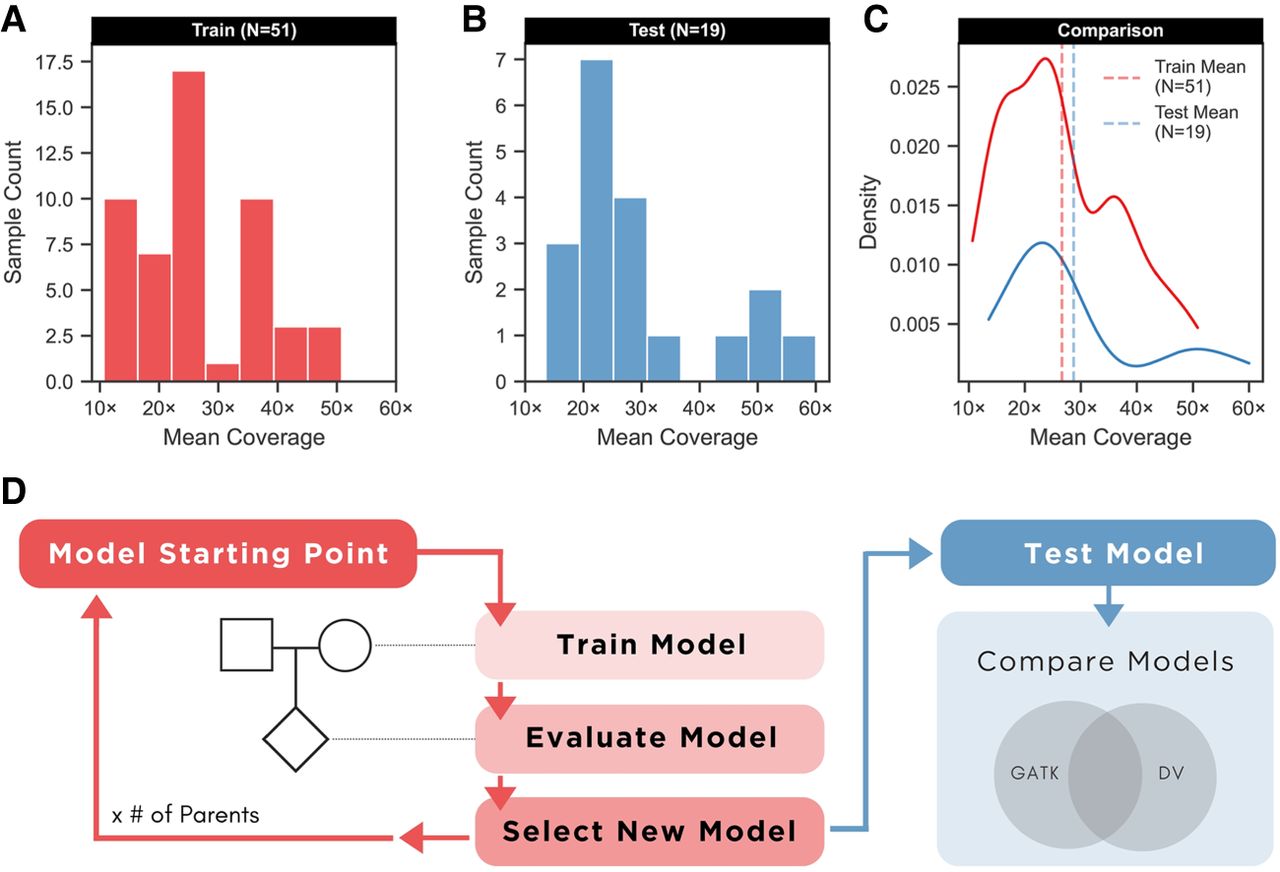
Before extending DeepVariant (DV), the available data were assigned to different purposes: training (A) or testing (B). The x-axis provides thresholds for genome-wide mean coverage, and the y-axis provides sample counts. (A) We selected samples from the UMAGv1 cohort with complete pedigree information (family trios). With TrioTrain, DV simultaneously uses labeled examples from two genomes: A parental genome provides new information, whereas learning is evaluated based on genotype predictions in the offspring. (B) Most samples used for model testing (13/19) lacked a complete pedigree but served as an independent test set because their GATK-based truth labels were withheld from DV during training. (C) The kernel density estimate (KDE) plot allows for comparing the two distributions after normalizing for sample size. (D) Workflow diagram of TrioTrain's iterative retraining process. All retraining begins with weights from an existing checkpoint (warm-starting); our experiments began with DV (v1.4) without the allele frequency channel. After initialization, all bovine-trained checkpoints include the allele frequency channel (DV-AF), encoding the PopVCF from the UMAGv1 cohort. Two iterations are performed for each trio, one for each parent. An iteration covers all labeled examples within a bovine parent–child duo constrained to sample-specific callable regions from Chr 1–29 and X. Each iteration produces a candidate checkpoint that is then assessed using a set of test genomes (N = 19). The resulting variant calls are compared against sample-specific, GATK-derived, truth labels to monitor training behavior over time.