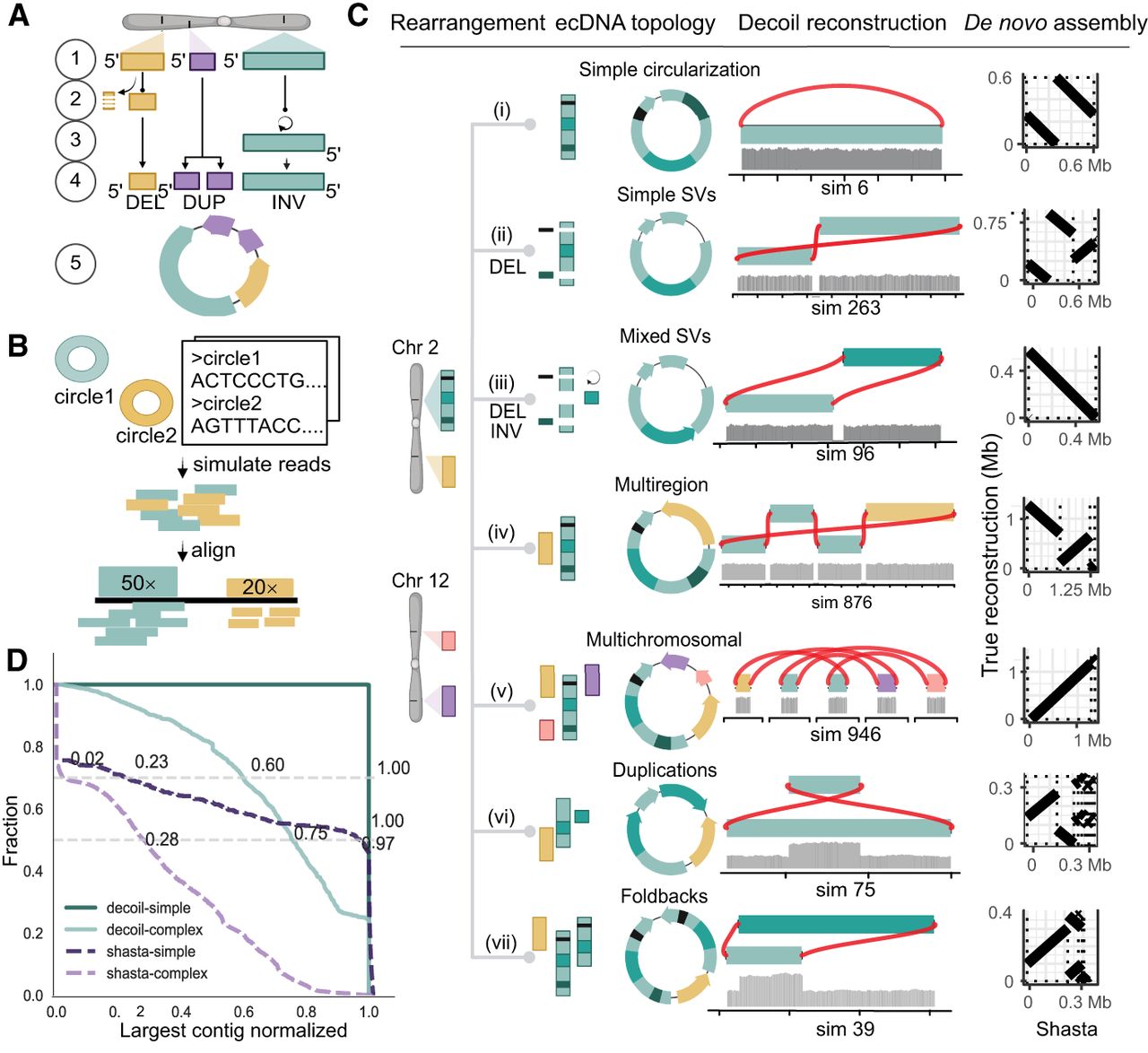
Decoil reconstructs complex ecDNA elements with high fidelity from simulated data. (A) Simulation strategy for generating individual ecDNA templates consists of the following steps: (1) choose genomic position, (2) simulate small deletions (DELs), (3) simulate inversion (INV), (4) simulate tandem-duplication (DUP), and (5) generate DNA sequence template (FASTA). The example depicts an ecDNA template harboring 1 × DEL (yellow), 1 × DUP (purple), and 1 × INV (green). (B) Pipeline for generating in silico long reads based on one or more ecDNA templates, at different depths of coverage. (C) The ecDNA topologies. Examples of ecDNA reconstructions performed by Decoil for simulated ecDNA elements, for the seven different topologies. The gray track represents the coverage of the aligned reads. The right column shows the Shasta de novo assembly (x-axis) against the true structure (y-axis). (D) Decoil and Shasta assembly contiguity for simple (i–v) and complex topologies (vi,vii). The y-axis represents the fraction of reconstructions with a specific contiguity (x-axis). The x-axis represents the larger contig normalized by the true structure length. One indicates a good reconstruction, zero poor reconstruction. Values greater than one refer to reconstructions larger than the true structure. The gray horizontal lines are at the 0.5 and 0.7 fractions.