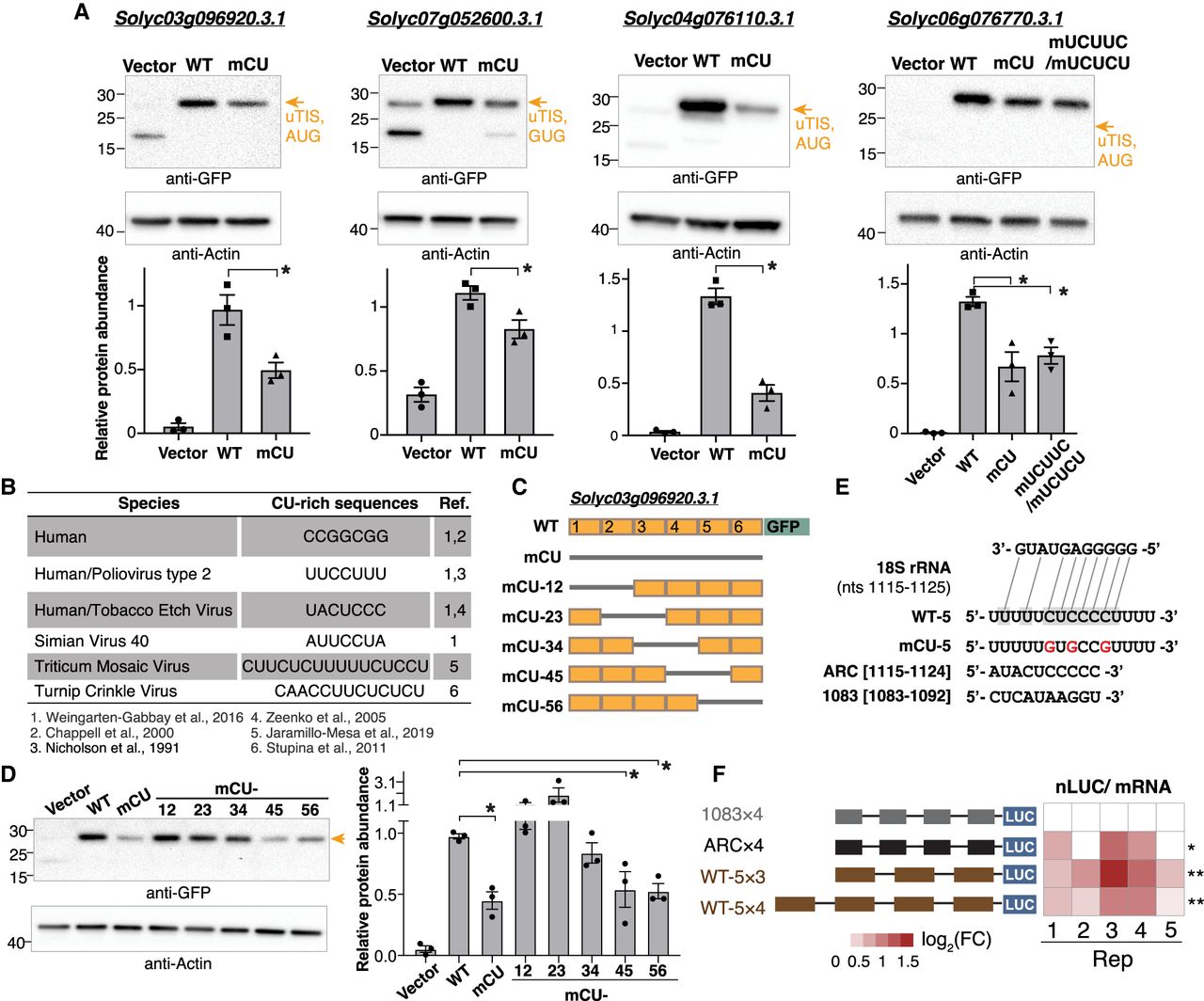
Mutation of CU-rich sequences on plant cellular mRNAs attenuated the TIS activities. (A) Immunoblotting analyses of proteins with translation initiated from the TISs indicated in Supplemental Figure S8 (orange arrows) and with translation driven by the upstream 100-nt wild-type (WT) sequence or sequences with mutations of CU tracts (mCU) or UCUUC/UCUCU (mUCUUC/mUCUCU) sites. Proteins were expressed in Nicotiana benthamiana (tobacco) leaves using the Agrobacterium-mediated transient expression system. Vector indicates tobacco leaves infiltrated with agrobacteria containing the expression vector (i.e., the GFP or FLAG-containing plasmid without a target gene sequence). (Bottom) The protein abundance of the reporter genes relative to actin for three biological repeats (dots) and the corresponding means and standard errors are shown. (B) The known CU-rich sequences found on human transcripts and on plant and animal viral mRNAs promote translation efficiency as reported in the indicated literature. (C) Illustration of swapping the WT (orange box) and the CU-mutated (mCU; gray line) sequences for the TIS-upstream region of a 5′ UTR–AUG TIS of Solyc03g096920.3.1 examined in A. The TIS-upstream region was divided into six subregions (indicated in Supplemental Fig. S8A) to generate distinct mCU mutants with indicated CU-mutated regions fused with the GFP reporter gene as described in A. (D) As described in A, but for the protein of the mCU mutants as indicated in C. (E) The putative binding site on WT sequences (shaded), but not on CU-mutated ones, to the plant 18S rRNA (sold and dashed lines). The sequences of the plant 18S rRNA at positions nt 1115 to 1125 and the WT sequence or sequence with mutations of CT tracts (red) in the fifth region indicated in C are shown. (*) P-values < 0.05, which is derived from one-way ANOVA test, representing the significant difference between WT and the samples with CU sequence mutations. (F) Heatmap shows the nLUC/ mRNA ratio for five biological replicates: 1083 × 4 and ARC × 4 denote negative and positive controls with four copies of 1083 and ARC fragments indicated in E. WT-5 × 3 and WT-5 × 4 denote vectors with three and four copies of the WT-5 indicated in E. (nLUC) Nano luciferase. The color keys represent log2 fold change (FC) relative to the negative control. (*) P-value < 0.05, (**) P-value < 0.01; determined by one-tailed Student's t-test.