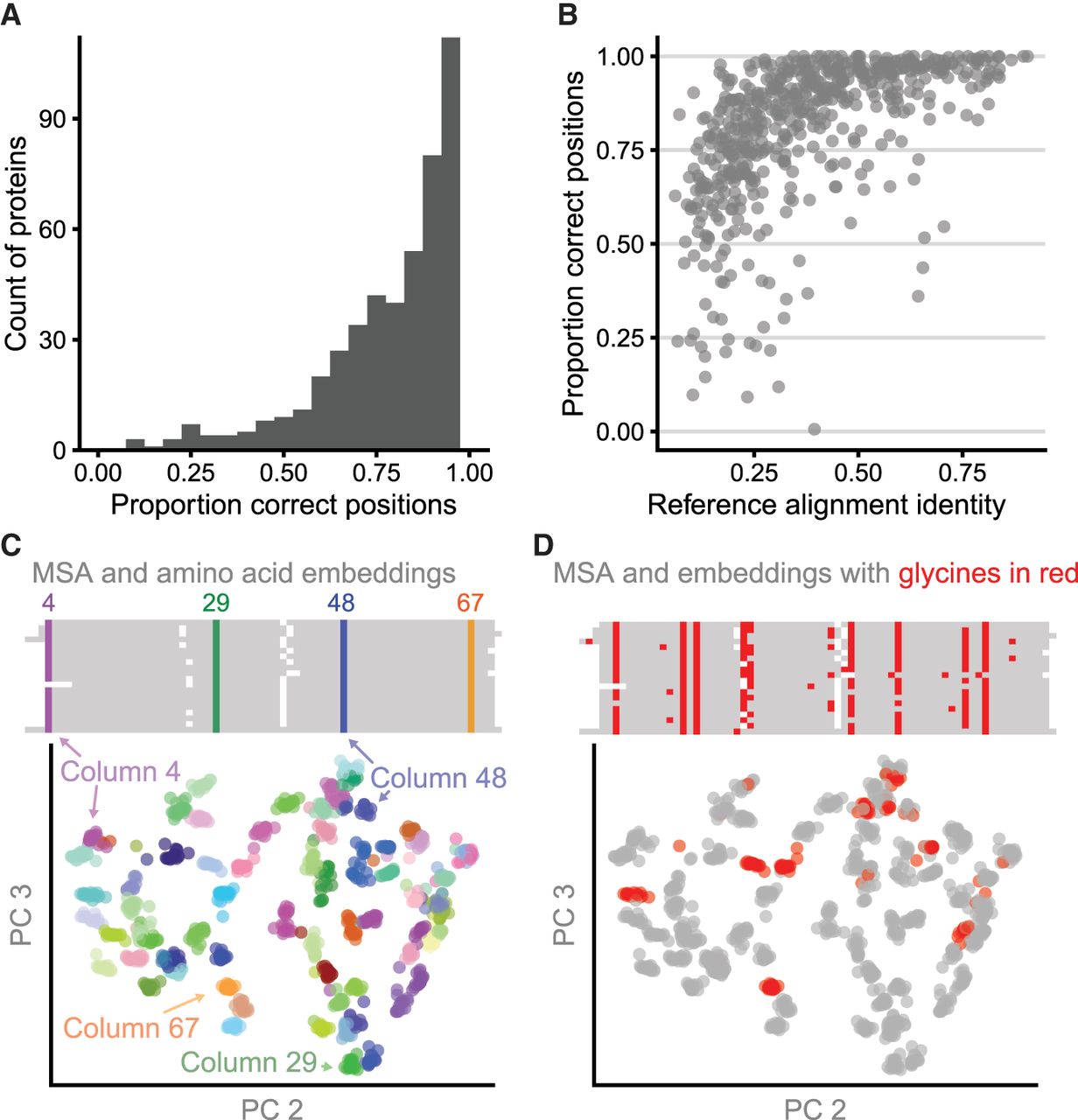
RBHs and clustered amino acid representations reflect aligned columns. (A) For 562 pairs of aligned proteins from the HOMSTRAD database, the MNCMs of RBHs between vector representations frequently correspond to correctly aligned positions. For each gold-standard alignment, we compute the fraction of aligned gold-standard positions that are additionally RBHs after MNCM filtering for that protein pair and display these results as a histogram. (B) The proportion of columns in the gold-standard pairwise alignments that are in the MNCM of RBHs is related to sequence identity of the reference alignment. (C) Principal components 2 and 3 for amino acid vector representations of 20 cold-shock proteins from the csp protein family. Amino acid representations (below) are colored by their corresponding column in the multiple sequence alignment (above). (D) Identical amino acids in different columns are distinguishable from each other. All glycines in the amino acid PCA plot and the multiple sequence alignment are colored red.