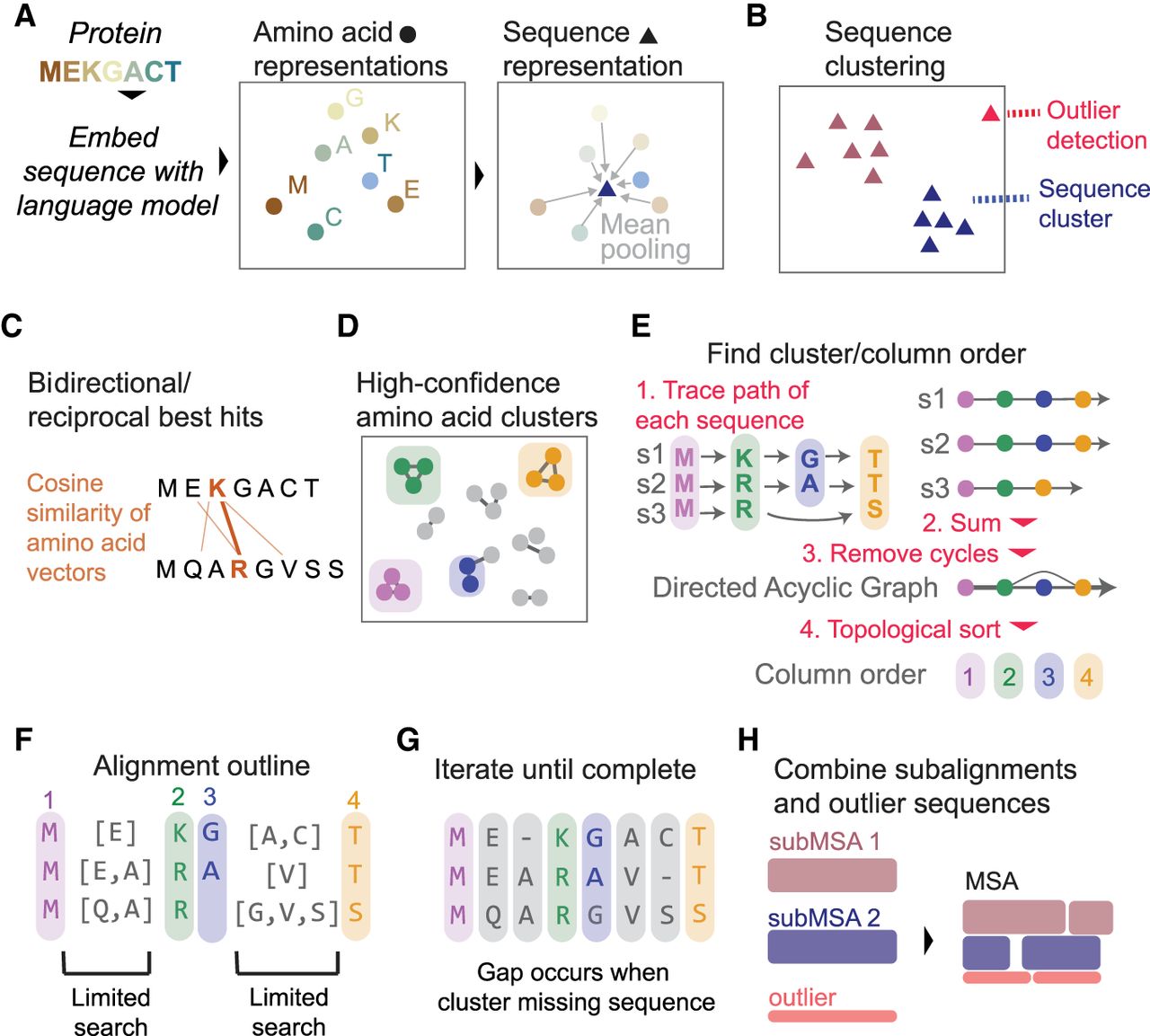
Overview of vcMSA algorithm. (A) Proteins are embedded using a protein language model to produce vector representations of each amino acid, and the mean of these amino acid embeddings is taken to produce a sequence-level representation. (B) We cluster sequence representations and detect outlier sequences. (C) For each sequence cluster, we determine bidirectional/reciprocal best hits (RBHs) of cosine similarity between pairs of amino acids in different sequences. (D) From a network built from RBHs, we determine confident clusters of amino acids, corresponding to columns in the MSA. (E) To determine column order, we trace the path of each sequence through clusters and combine all paths into one network, taking edge weights from the number of sequences that traverse between the pairs of clusters. We trim any clusters that cause cycles and use a topological sort of the resulting directed acyclic graph to find column order. (F) Clusters/columns limit scope of search for unplaced amino acids. (G) We iterate limited searches until all amino acids are placed. Gaps in the alignment occur when a cluster does not contain an amino acid from a sequence. (H) We combine alignments from each sequence cluster and outliers in the final output MSA.