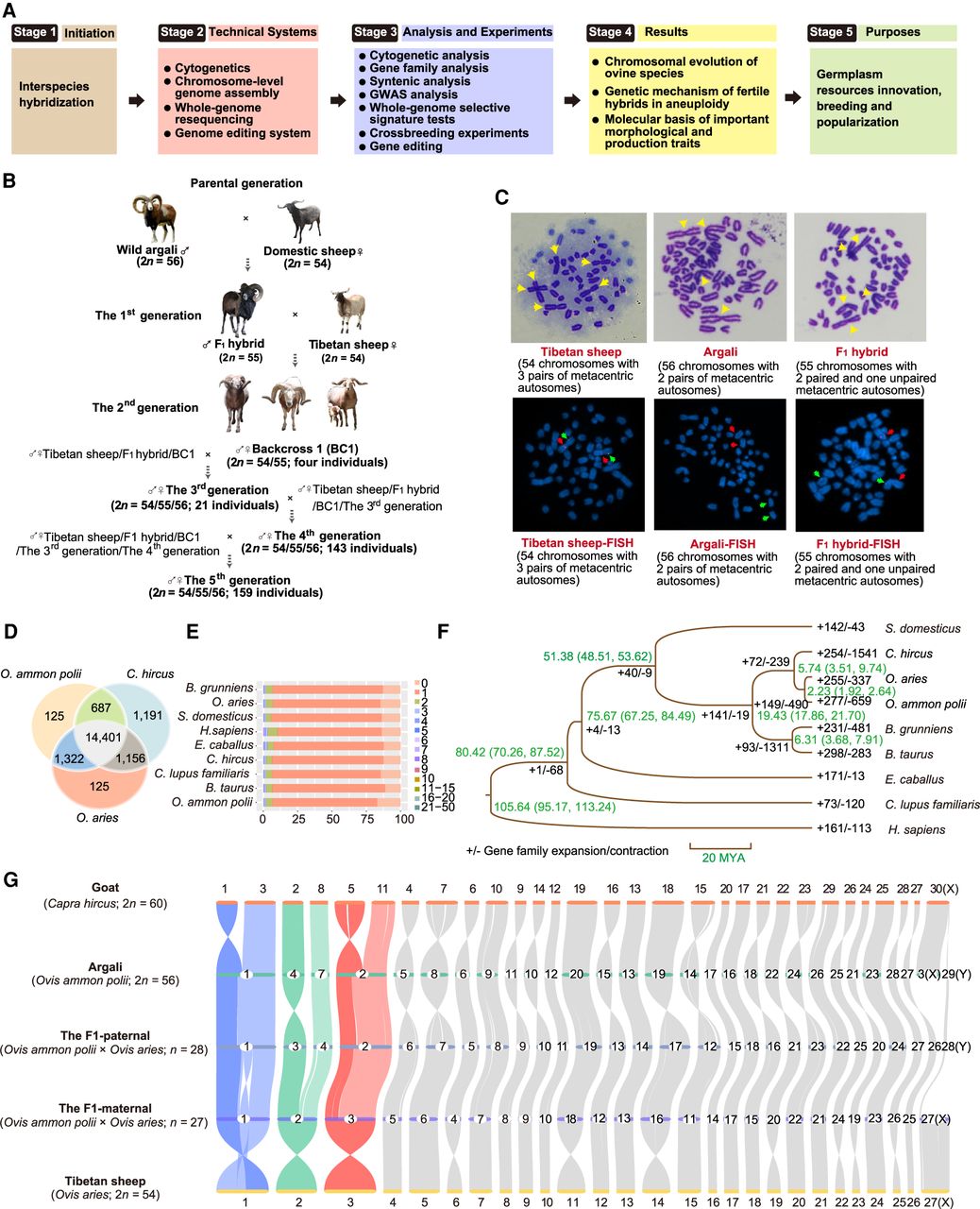
Research framework, hybridization scheme, karyotype and chromosomal painting, statistics of homologs and gene families, phylogenetic tree, and synteny landscape. (A) Roadmap of initiation, technical systems, analysis and experiments, the results, and purposes of this study. (B) The wild (argali, O. ammon)-domestic sheep hybridization scheme. The F1-hybrid is subsequently backcrossed to Tibetan sheep to produce the progenies, and later hybrid generations are produced by intercrossing and backcrossing. (C) Karyotype and chromosomal painting of Tibetan sheep, argali, and the F1-hybrid. DNA probes (GNAQ: green signal; STK39: red signal) were used to label the long arm and short arm of Chromosome 2 and two acrocentric chromosomes. DNA was stained with 4,6-diamidino-2-phenylindole (DAPI; blue). (D) Number of homologs among the Marco Polo subspecies of argali (O. ammon polii), sheep (O. aries), and goat (Capra hircus). (E) Proportions of different gene numbers in 19,917 gene families in the nine mammal species. (F) Phylogenetic tree of nine mammal species based on 10,043 single-copy orthologous genes. Numbers in green to the right of nodes are the divergence times and their 95% confidence intervals (95% CIs). Values next to the branches represent the numbers of gene family expansions/contractions. (MYA) Million years ago. (G) Genome-wide syntenic relationship among goat (C. hircus), sheep (O. aries), the Marco Polo subspecies of argali (O. ammon polii), and the argali-domestic sheep F1-hybrid. Syntenic blocks involved in chromosome fusion are marked with colored ribbons.