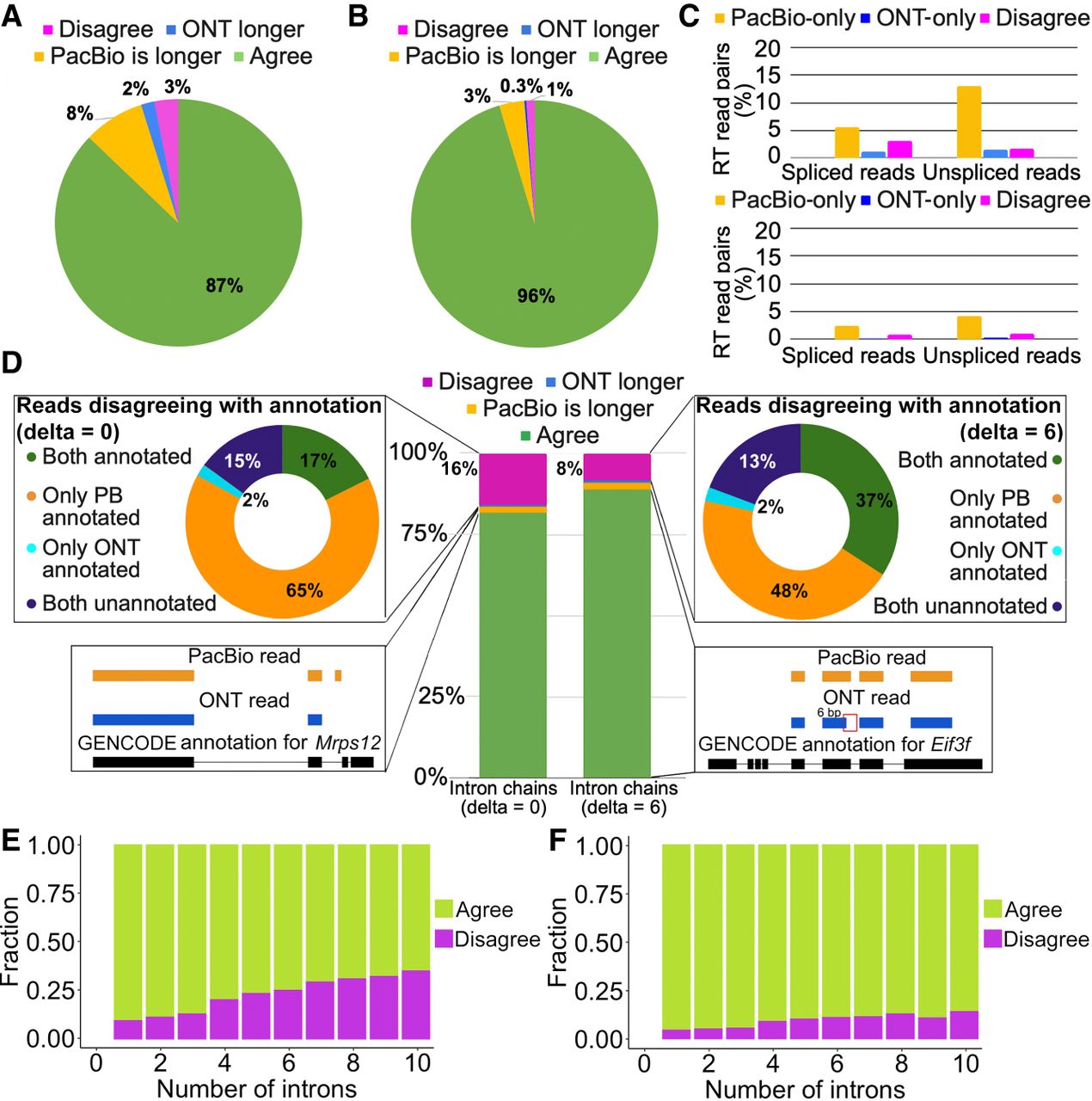
Agreement in RT read pairs of Sl-ISO-Seq data. (A) Fractions of TSS assignments that agree (green), disagree (magenta), or are found only in one read (blue for ONT, yellow for PacBio) from the RT read pair. (B) Same as A, but for poly(A) sites. (C) Percentage of RT read pairs that disagree on the assigned TSS (top) and poly(A) site (bottom): only PacBio read assigned (yellow), only ONT (blue), and both assigned but to different sites (magenta). (D, middle) Percentage of RT read pairs that agree (green), disagree (magenta), or have one chain being longer (blue for ONT, yellow for PacBio [PB]) when splice junctions are compared precisely (left; delta = 0bp) or inexactly (right; delta = 6bp). (Top left) Classification of disagreeing intron chains from RT read pairs with respect to the reference annotation (delta = 0 bp): Both are inconsistent with the annotation (dark blue); both correspond to known (different) transcripts despite the disagreement (green); and PacBio is consistent with the annotation whereas ONT is not (yellow) and vice versa (light blue). (Top right) Classification of disagreeing intron chains with respect to the reference annotation using inexact comparison (delta = 6 bp). (Bottom left) An example of agreeing intron chains from an RT read pair in which PacBio intron chain is longer (Mrps12 gene). (Bottom right) An example of intron chains from an RT read pair that have a 6-bp difference in the donor site of the second intron. Comparing intron chains with delta = 6 bp classifies them as agreeing (Eif3f gene). (E) Fraction of agreeing (green) and disagreeing (magenta) intron chains with respect to intron chain length when compared precisely (delta = 0 bp). (F) Same as E, but with delta = 6 bp.