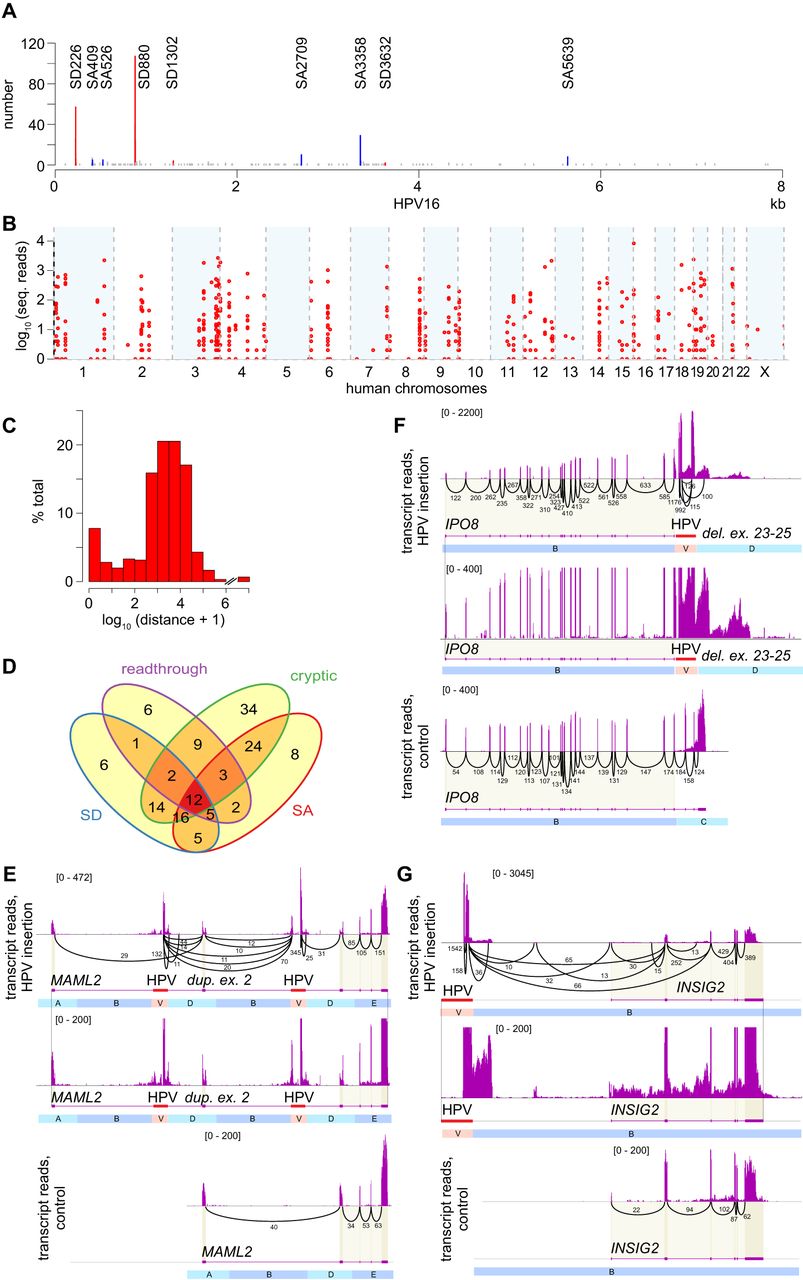
HPV integrants induce various forms of genetic disruption including gene breakage and chimeric transcription. (A) Counts of virus–host chimeric transcript junctions (y-axis) in 91 HPV16-positive tumors, aligned to the HPV16 genome (x-axis) with known splice donor (SD coordinate, red), splice acceptor (SA coordinate, blue), and other (gray) sites. (B) Counts of split or discordant RNA-seq reads (y-axis) in 103 HPV-positive tumors supporting chimeric transcript junctions (n = 673), aligned to the human genome (x-axis). (C) Frequency distribution (y-axis, percent total) of log10-transformed genomic distances (x-axis) between virus–host junctions from RNA-seq versus nearest DNA breakpoint (n = 604). (D) Venn diagram counts chimeric transcripts expressed at 147 genes, via host splice donor (SD, blue, n = 61); splice acceptor (SA, red, n = 75); readthrough transcription (purple, n = 40), and/or cryptic splice sites (green, n = 114). (E–G) Sashimi plots depict counts of mapped RNA-seq reads at genes with HPV integrants in affected tumor (top, center panels) versus without integrants in control tumor (bottom). Intron sequences not shown to scale. Center, bottom panels, identical scale of reads (y-axis, brackets). Black arcs, numbers, read counts connecting spliced exons. (E) Intragenic HPV integrants in MAML2 (red) flank a ∼75-kb duplication including exon 2, and delete small intronic segment C. Gene breaking involves premature transcriptional termination of MAML2 after exon 2 and de novo initiation of downstream transcripts from HPV. Segment B is truncated for visualization. (F) Intragenic HPV integrants in IPO8 (red) delete distal exons 23–25, disrupting 3′ transcripts and up-regulating upstream exons. (G) Intergenic HPV integrants both upstream of and downstream from INSIG2 flank a ∼665-kb duplication on Chr 2q14, extending from Chr 2:118.826 to 119.492 Mbp. Numerous chimeric transcripts originating from an upstream, intergenic HPV16 integrant are spliced to a novel exon, novel splice acceptor site, and exons 1, 2, and 3, causing gene disruption. See also Supplemental Figures S5.1–S5.4 and Supplemental Tables S6.1–S6.6.