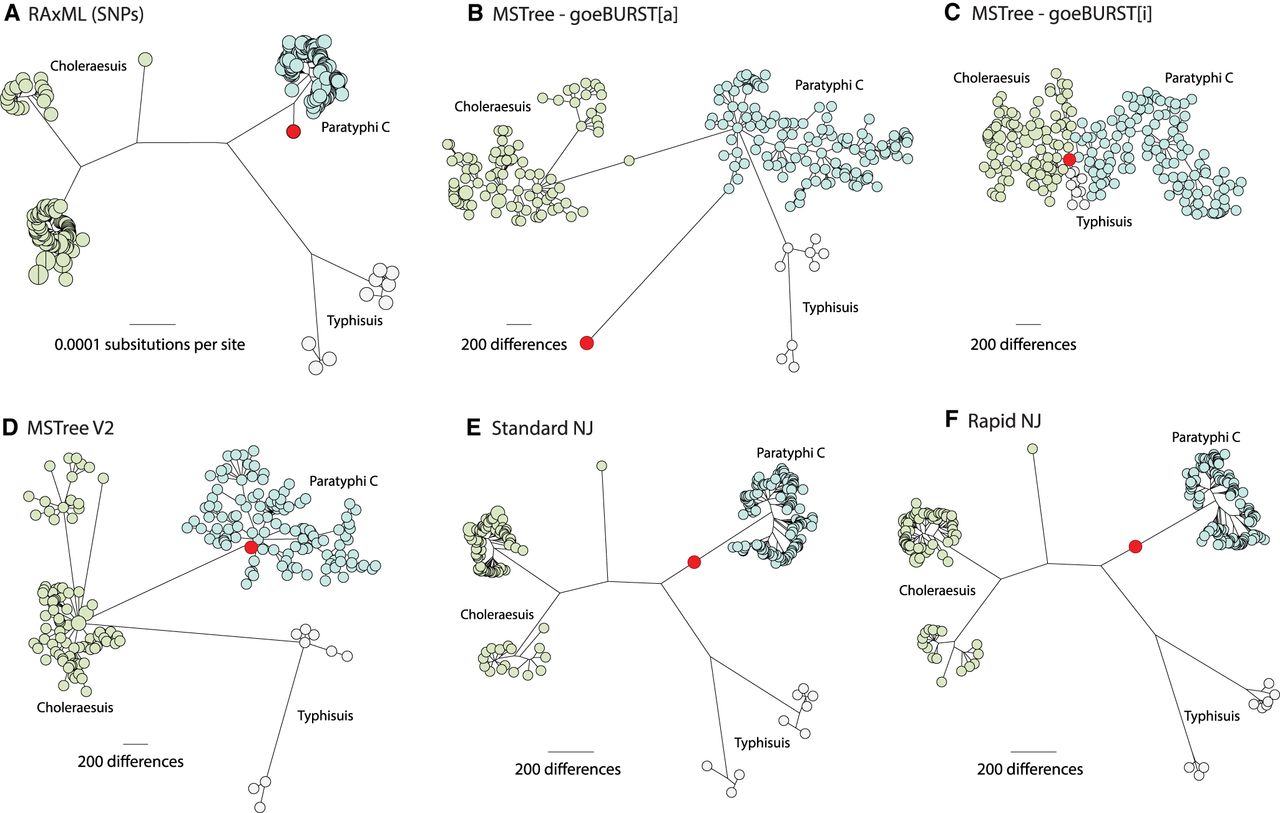
Comparisons of different topologies produced by six algorithms when extensive missing data is present. Trees were calculated from 20,114 nonrecombinant, core genomic SNPs (A) or 3002 loci in the cgMLST V2 scheme (Alikhan et al. 2018) (B–F) that were found in 218 modern genomes from Salmonella serovars Paratyphi C, Typhisuis, or Choleraesuis (Zhou et al. 2018). The modern genomes were supplemented by one ancient genome (Ragna; red) that had been reconstructed from an 800-yr-old skeleton. The algorithms used were maximum likelihood (A, RAxML), MSTree (B, GoeBURST[a]; C, GoeBURST[i]), MSTree V2 (D, GrapeTree), NJ (E, FastMe), or RapidNJ (F, RapidNJ), and all trees were visualized in GrapeTree SA. Nodes are color-coded by serovar. Due to fragmentation in the ancient Ragna DNA and intermediate levels of genome coverage, cgMLST alleles in Ragna could only be called for 215 (<10%) of the 3002 cgMLST loci, and the remainder of the cgMLST alleles were scored as missing data. Similarly, only 19,245 (96%) of the SNPs could be called in the Ragna genome. The Ragna genome is on a side-branch that diverged prior to the coalescence of the crown branch leading to modern Paratyphi C and differs from that coalescent by 263 SNPs. The correct position and branch length of the Ragna branches are as shown in A. Ragna is on an artificial, long terminal branch in B because all missing data count as different alleles. Ragna is central in part C because it is ≤215 cgMLST allele differences from all modern genomes and therefore forms an artificial central hub for all the genomes. MSTree V2 (D) maps Ragna to a tiny side branch preceding the Paratyphi C coalescent, similar in topology to how it was mapped on the basis of SNPs (A). However, NJ (E) and RapidNJ (F) mapped Ragna incorrectly near the base of the long, main branch leading to the crown group of Paratyphi C. The mapping of Ragna to the main branch rather than on its own side branch resulted because those algorithms calculated a negative distance from Ragna to the main branch. Interactive versions of each tree are available at (A) http://bit.ly/2vuFIIb, (B) http://bit.ly/2HF5tYt, (C) http://bit.ly/2qDD3GT, (D) http://bit.ly/2JRBvkQ, (E) https://bit.ly/2B6IS7v, and (F) https://bit.ly/2z2LWRb.