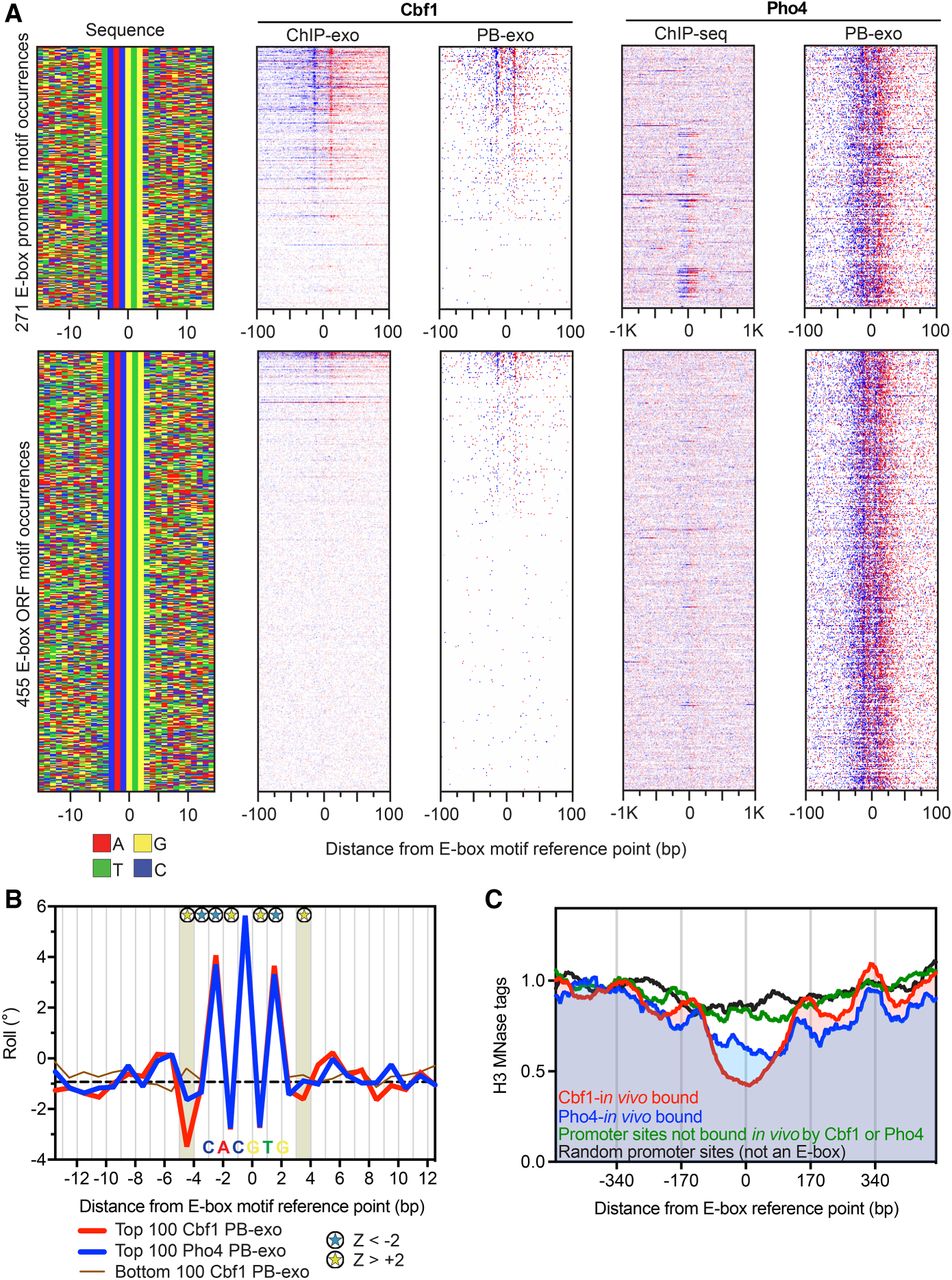
Genome-wide in vitro Cbf1 and Pho4 binding locations. (A) Annotation descriptions are the same as in Figure 2A, except for Cbf1 and Pho4. Data are sorted by Cbf1 PB-exo tag counts ±30 bp from the motif center, and rows across all data sets are linked. The Pho4 ChIP-seq data under phosphate starvation is from Zhou and O'Shea (2011). (B) Line plots of variations in roll for the top (red) versus top (blue) 100 Pho4 PB-exo-bound E-box motif occurrences. The bottom (brown) 100 Cbf1 PB-exo motif occurrences are also shown but were not included in the statistical analysis. The dashed black line indicates the genome-wide median. Blue and yellow stars represent positions with significant large or small roll (|Z| > 2, Mann-Whitney U test), respectively. The shaded area designates the positions just outside the core motif that possessed significant differences in DNA shape. The position of the consensus E-box motif is labeled along the x-axis. (C) Composite plots of nucleosome dyads generated by MNase H3 ChIP-seq for different groups of E-box motif occurrences. The data were collected from cells grown in YPD, but the Pho4-in vivo bound sites were defined by data collected under phosphate starvation conditions.