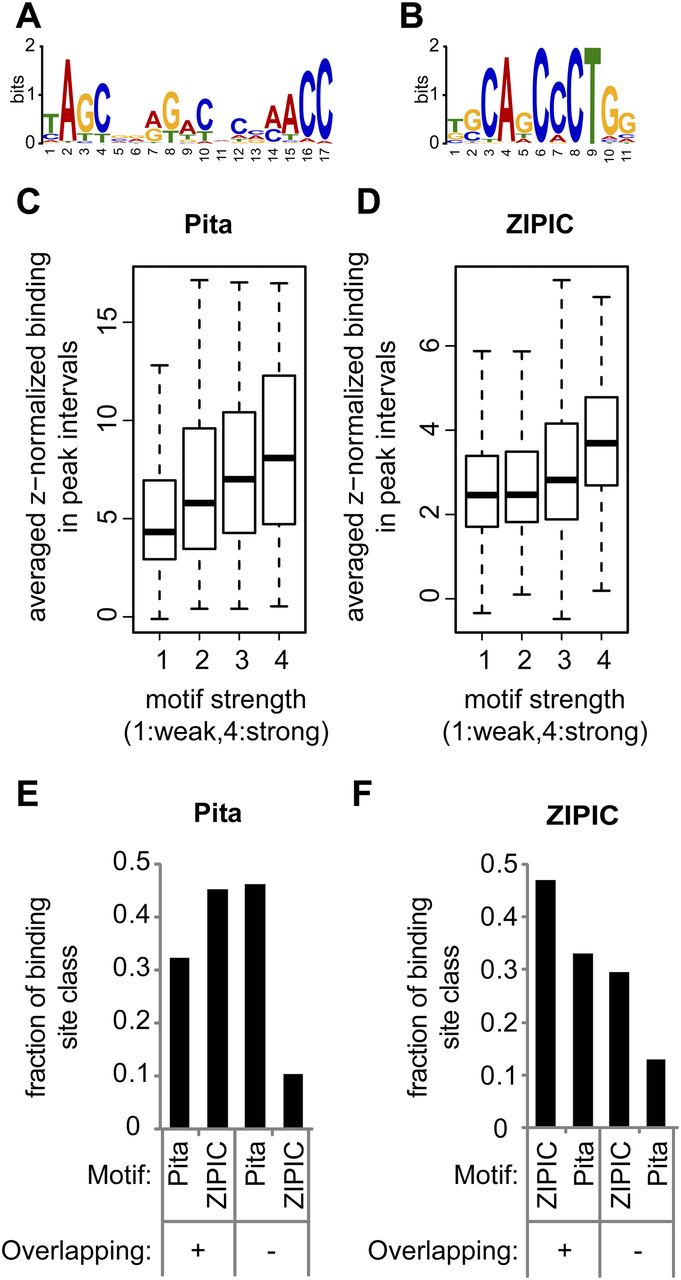
Figure 4.
Motif specificity for Pita and ZIPIC. (A,B) The top motifs identified within (A) Pita and (B) ZIPIC peaks as a sequence logo. (C,D) The binding of (C) Pita and (D) ZIPIC is correlated with motif conservation. Peaks were classified with respect to similarity to the best matching consensus-like sequence and grouped accordingly: from (1) weak similarity to (4) high similarity. (E,F) Peaks of (E) Pita and (F) ZIPIC were divided into two groups by the criterion of overlap with peaks of the other protein (overlapping versus nonoverlapping peaks), and both groups were analyzed for the presence of ZIPIC and Pita motifs.