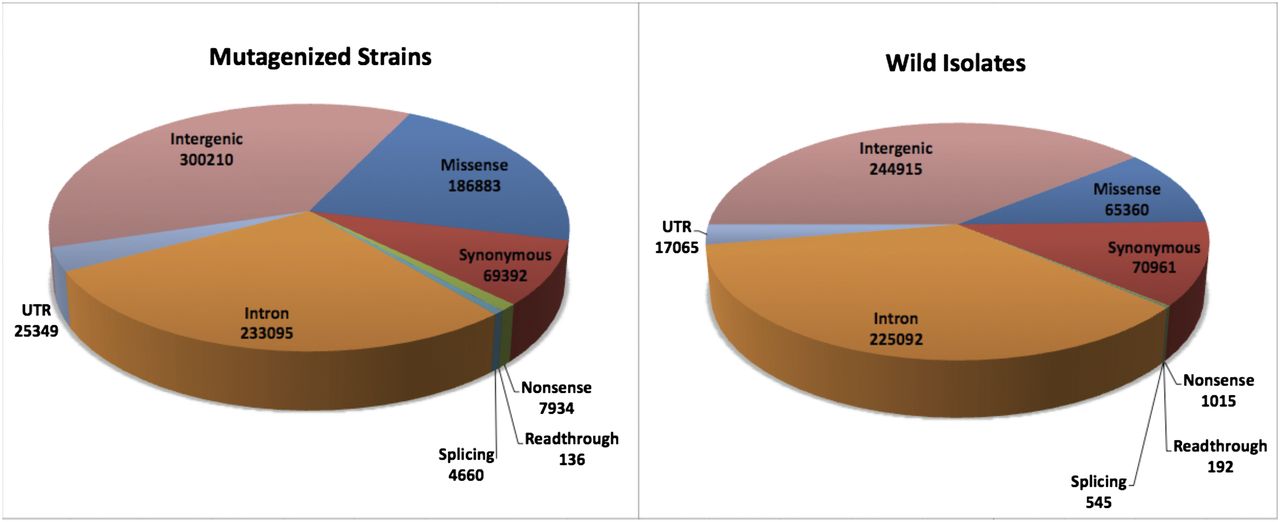
Figure 4.
Mutation effects (SNVs) in mutant strains and wild isolates. The inferred effects of the SNVs in the mutant strains and wild isolates are plotted, showing the disparity in the fraction of mutations affecting coding sequence. Mutations resulting in nonsynonymous, nonsense, and splice site changes are three to 10-fold less frequent in the wild isolates, whereas synonymous changes are similar in number between the two. In addition, the number of mutations in intronic regions is similar between mutant and WI collections, but both UTR and intergenic sites are less abundant in the WI strains.