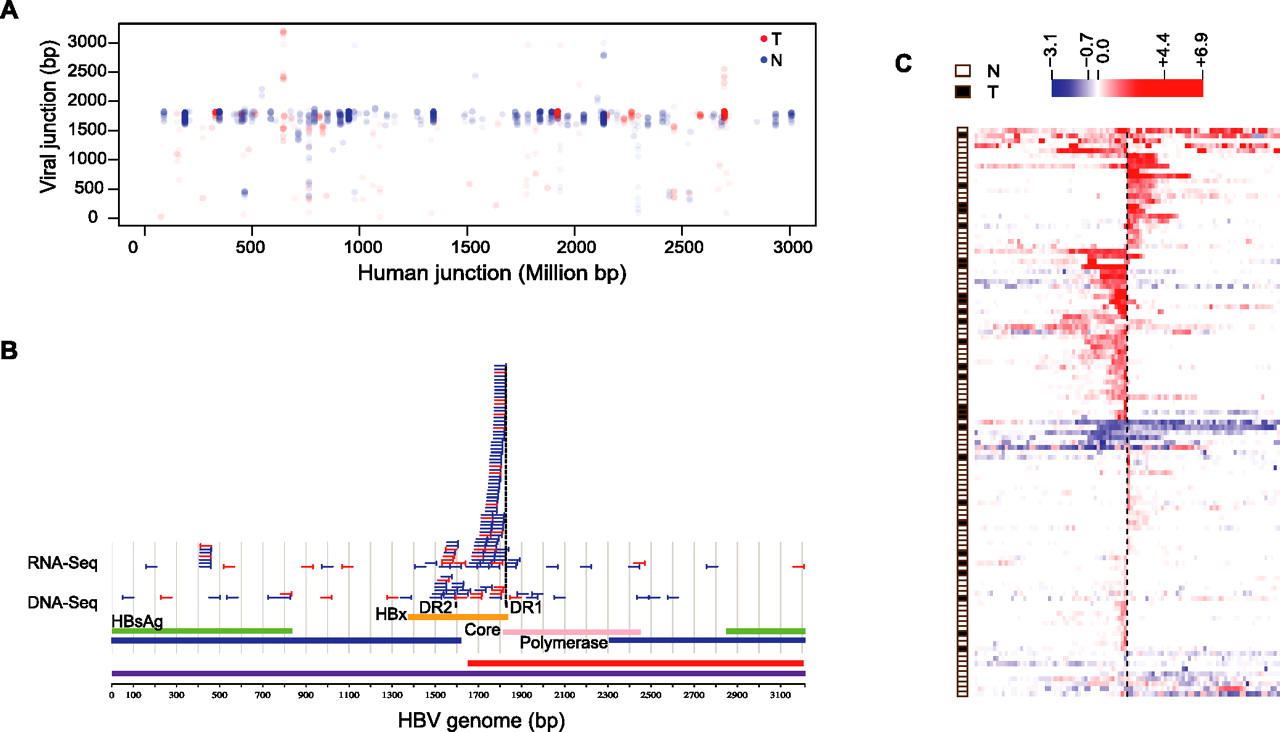
Viral-human fusion transcripts are common in both HCC and nontumor samples. (A) The genomic coordinate on the HBV genome of each chimeric RNA-seq read is plotted against its genomic coordinate on the human genome (linearized after concatenating all chromosomes). Only locations supported by two or more chimeric reads are shown. (B) Viral junctions determined from clusters of two or more chimeric reads are shown as vertical bars, with part of the remaining integrated viral sequence (50 bp) indicated as horizontal lines. Reads from both RNA- and DNA-seq are shown. A large majority of the RNA-seq junctions are in close proximity (10 bp) to the DR1 (Direct repeat 1) region on the HBV genome. (Blue) Clusters from nontumor liver samples; (red) clusters from tumor samples. (C) A global view of the transcriptional consequence of viral integration on the flanking human genome. The data were organized in the same manner as in Figure 2D, except that the human junctions in this panel are based on RNA-seq data instead of DNA-seq data. Most of the sites show strong unidirectional transcriptional up-regulation, starting at the integration site, while relatively fewer sites correlate with nondirectional transcriptional down-regulation or up-regulation.