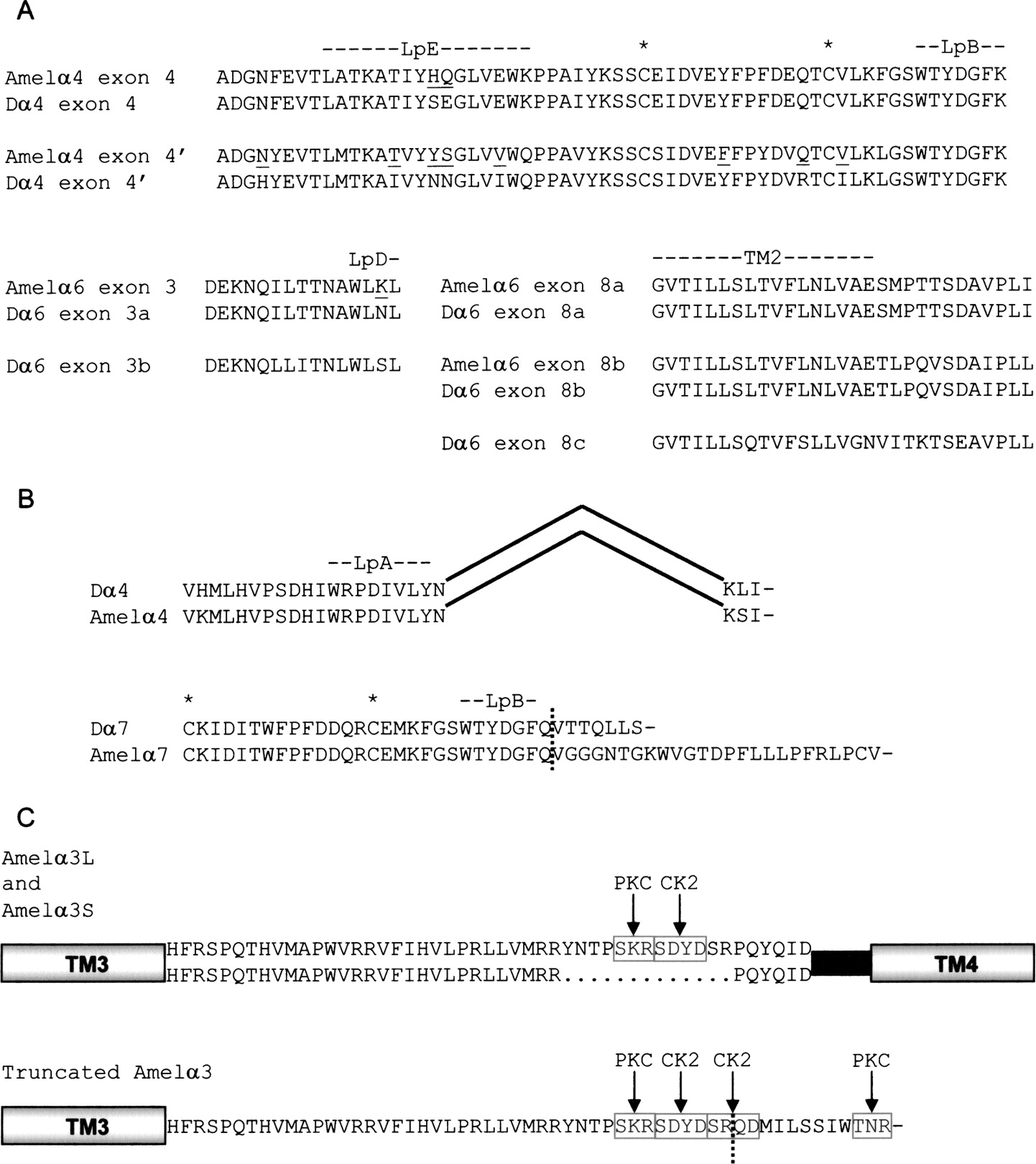
Splice variants of A. mellifera nAChR subunits. (A) Comparison of alternative exons of D. melanogaster and A. mellifera. Apis residues that differ from those of the orthologous Drosophila exon are underlined. (B) Conservation of truncated nAChR transcripts between D. melanogaster and A. mellifera. Omission of exon 4 of Dα4 and Amelα4 results in a frameshift and premature termination of translation, while nonsplicing of intron 5 of Dα7 and the equivalent intron of Amelα7 introduces a loss of the reading frame and a premature stop codon. (C) Novel splice variants in Apis nAChRs. The TM3 and TM4 domains are represented schematically, while the sequence of a portion of the intervening intracellular loop is shown highlighting the three splice variants, and the remainder of the loop is represented by a black bar. Loops involved in ligand binding and transmembrane regions are indicated, while the cys-loop is marked by asterisks. Potential phosphorylation sites are highlighted by gray shading and broken lines mark the start of altered reading frames resulting from unspliced introns.