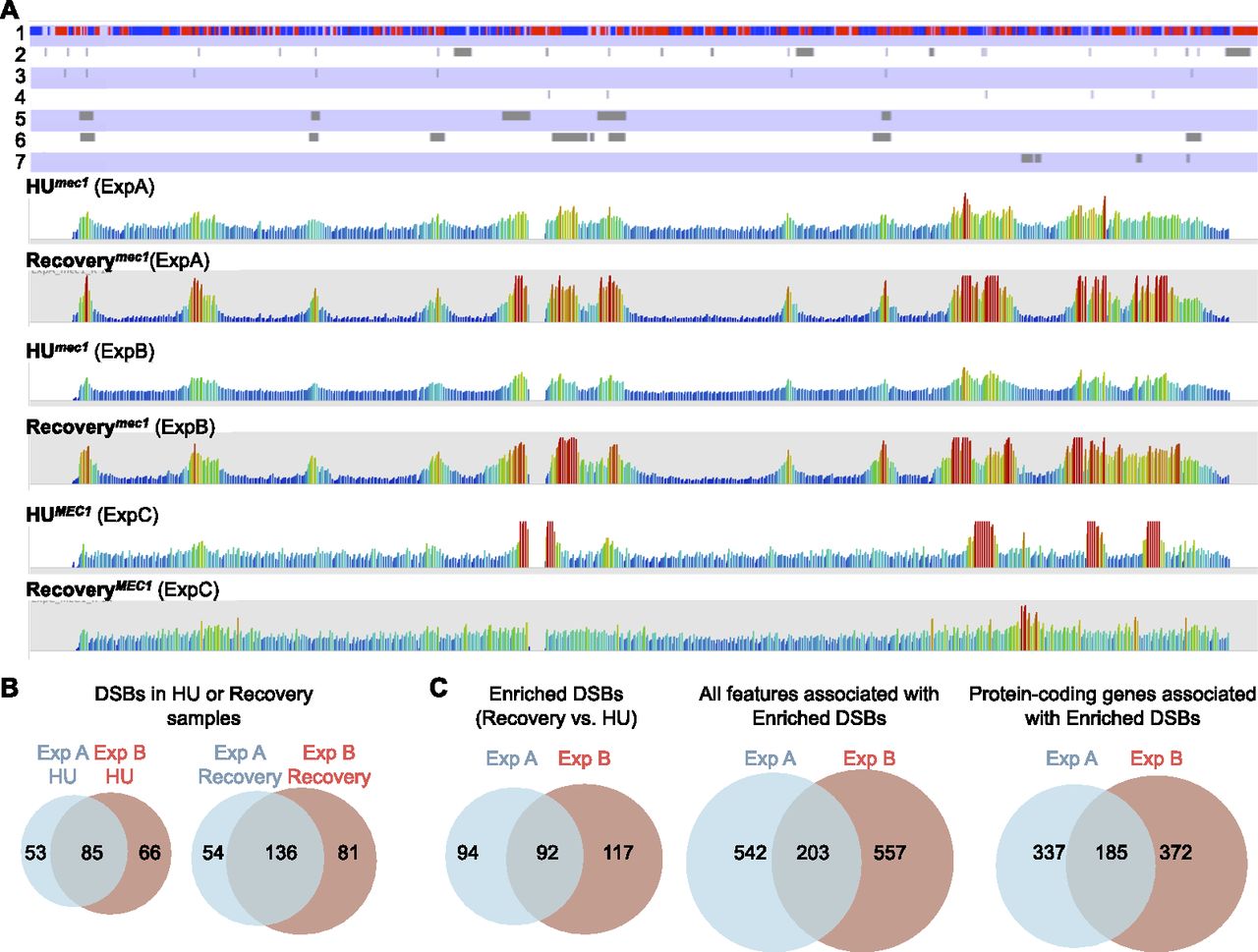
Chromosome views of the Break-seq data using SeqMonk. Sequence reads quantification was performed as described in Supplemental Figure S1. (A) View of Chromosome X (745,751 bp) of the HU and Recovery samples from all experiments. The annotation tracks from top to bottom are (1) ORFs (red, Watson-strand-encoded; blue, Crick-strand-encoded); (2) OriDB-curated confirmed and likely ARSs; (3) Rad53-checked origins (Feng et al. 2006); (4) Rad53-unchecked origins (Feng et al. 2006); (5) enriched DSBs in Exp A; (6) enriched DSBs in Exp B; and (7) enriched DSBs in Exp C. The data tracks are as labeled. (B) Venn diagrams showing concordance between the DSBs in HU samples alone and between the DSBs in Recovery samples alone in the two experiments. (C) Venn diagrams showing the concordance between the enriched DSBs in two experiments (left) and the identification of 203 common features (middle) and 185 common protein-coding genes (right).