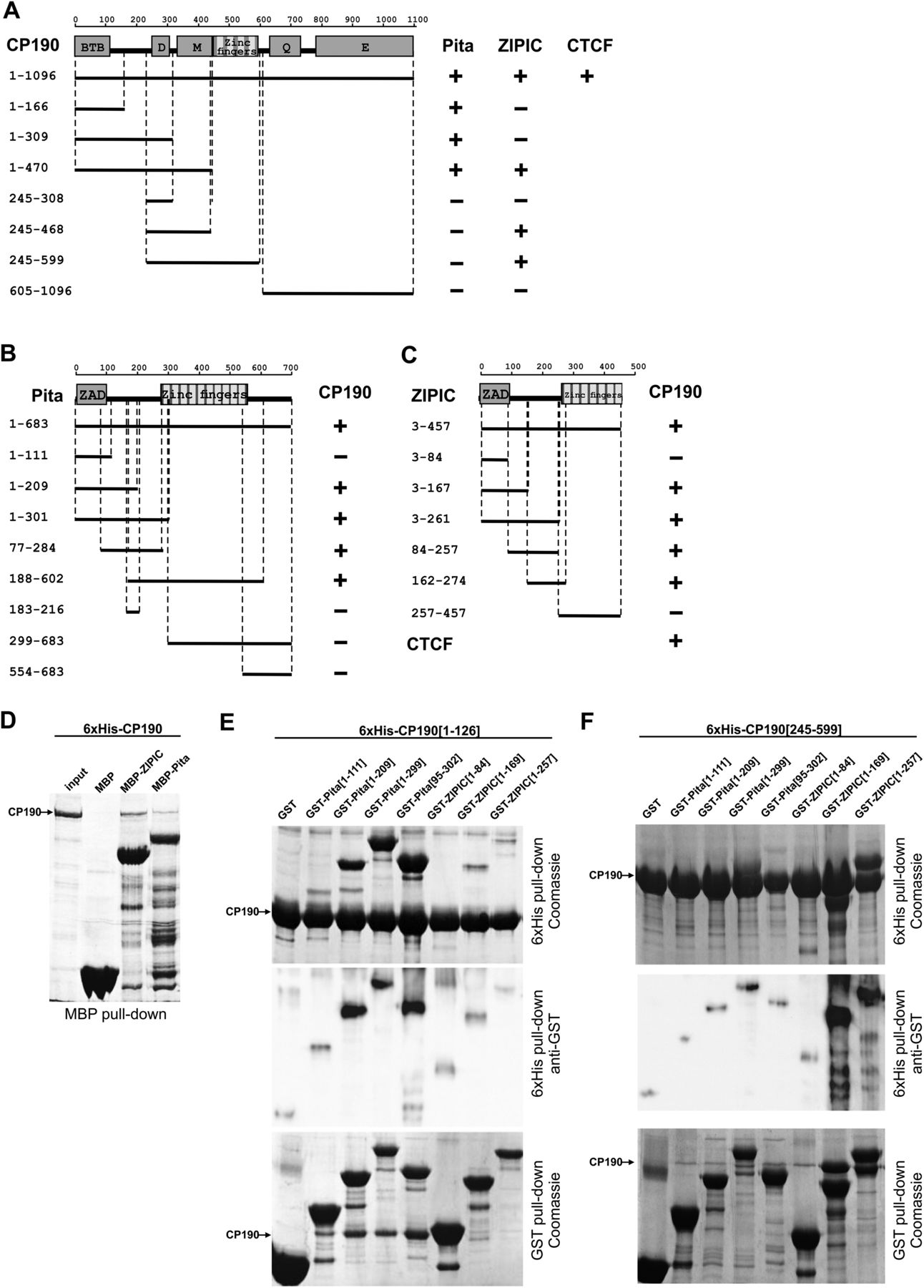
Identification of interacting domains of CP190, Pita, and ZIPIC proteins. (A) Localization of CP190 domains interacting with Pita and ZIPIC in yeast two-hybrid assay. In the structural scheme of full-length CP190, protein domains are shown as boxes, and lines indicate the different deletion fragments. The horizontal scale and figures on the left show the positions of amino acid residues. Different fragments of CP190 were fused to the GAL4 DNA-binding domain and tested for interaction with Pita and ZIPIC fused to the GAL4 activating domain. All CP190 fragments were tested for the absence of interaction with the GAL4 activating domain alone, whereas Pita and ZIPIC were tested for the absence of interaction with GAL4 DNA-binding domain alone. The results are summarized in columns on the right, with the “+” and “−” signs referring to a strong interaction or the absence of interaction, respectively. Interaction of CP190 with dCTCF was used as a positive control. (B) Localization of Pita domains interacting with CP190 in a yeast two-hybrid assay. Different fragments of Pita were fused to the GAL4 activating domain and tested for interaction with CP190 fused to the GAL4 DNA-binding domain. All Pita fragments were tested for the absence of interaction with the GAL4 DNA-binding domain alone. Other designations are the same as in A. (C) Localization of ZIPIC domains interacting with CP190 in a yeast two-hybrid assay. Different fragments of CG7928 were fused to the GAL4 activating domain and tested for interaction with CP190 fused to the GAL4 DNA-binding domain. All ZIPIC fragments were tested for the absence of interaction with the GAL4 DNA-binding domain alone. Other designations are the same as in A. (D) Interaction of full-length recombinant CP190 and Pita/ ZIPIC in MBP pull-down assay. Beads with bound MBP-Pita, MBP-ZIPIC, or MBP alone were incubated with 6 × His-CP190, and the precipitated proteins were resolved by SDS-PAGE and stained with Coomassie. (E,F) Interaction of CP190 fragments [1–126] (E) and [245–599] (F) with different fragments of Pita and ZIPIC in GST and 6 × His pull-down assays. The positions of amino acids are indicated by square brackets. In the 6 × His pull-down assay, beads with bound 6 × His-CP190 were incubated with each GST-tagged Pita or ZIPIC fragment or GST alone. In the GST pull-down assay, beads carrying GST-tagged Pita or ZIPIC fragment or GST alone were incubated with 6 × His-CP190 fragments. The precipitated proteins were resolved by SDS-PAGE and stained with Coomassie. Additionally, the precipitated proteins were immunoblotted with anti-GST antibodies in the case of the 6 × His pull-down assay.