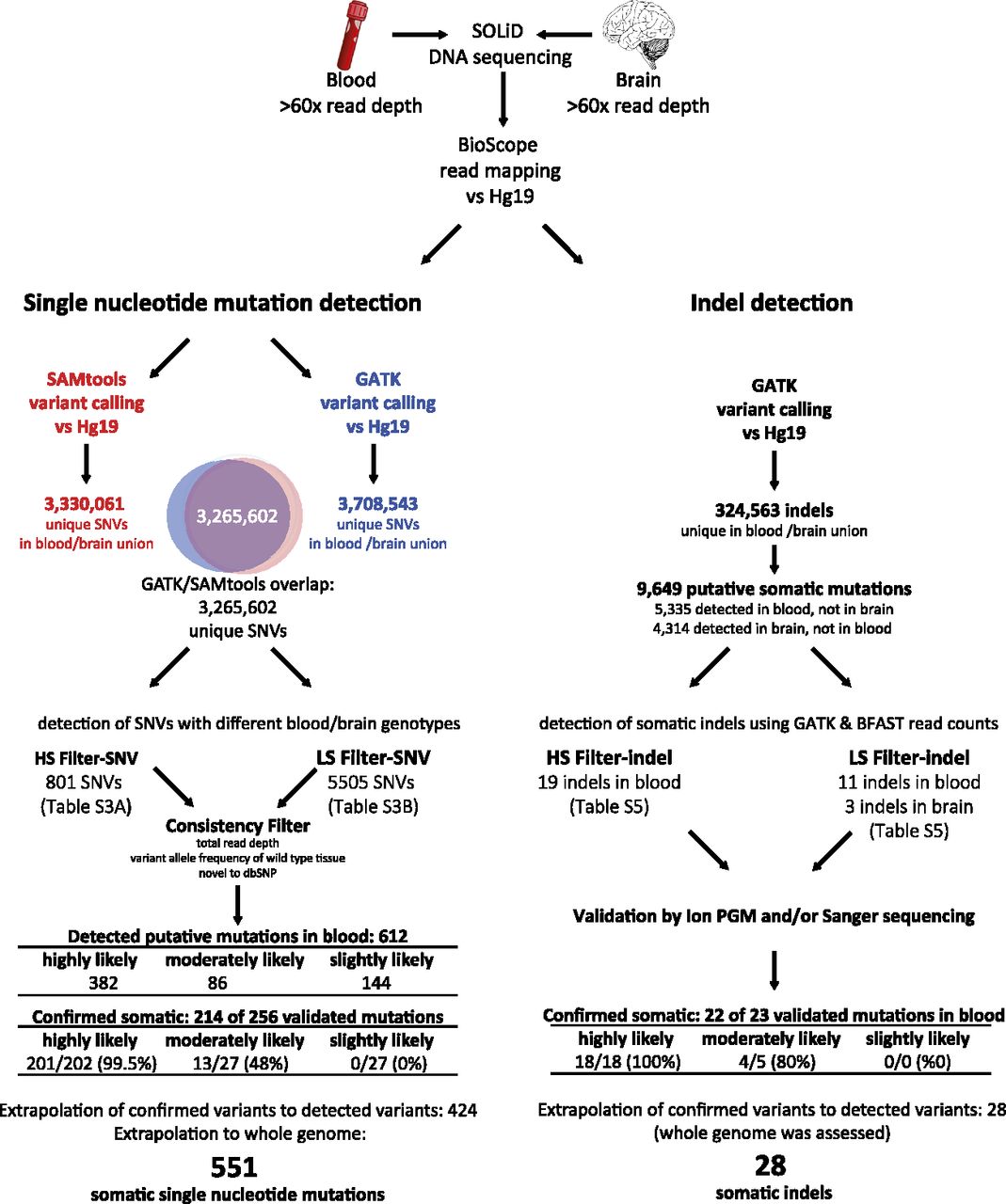
Somatic variant detection pipeline SNV and indel detection. Whole-genome sequences of W115 blood and brain tissues were mapped to hg19. Variants were called using both GATK and SAMtools. The SNVs that overlapped between the two genotyping algorithms were considered most trustworthy and were used for further analysis with high stringency (HS-SNV) and low stringency (LS-SNV) filters. Indels were passed through a high stringency (HS-indel) and low stringency (LS-indel) read-depth filter. Candidate somatic SNVs and indels were confirmed by validation with Ion Torrent PGM sequencing and/or Sanger sequencing. The number of confirmed SNVs and indels was extrapolated to account for those not validated. The number of SNVs was also extrapolated to the whole genome, whereas for indel detection the whole genome was assessed.