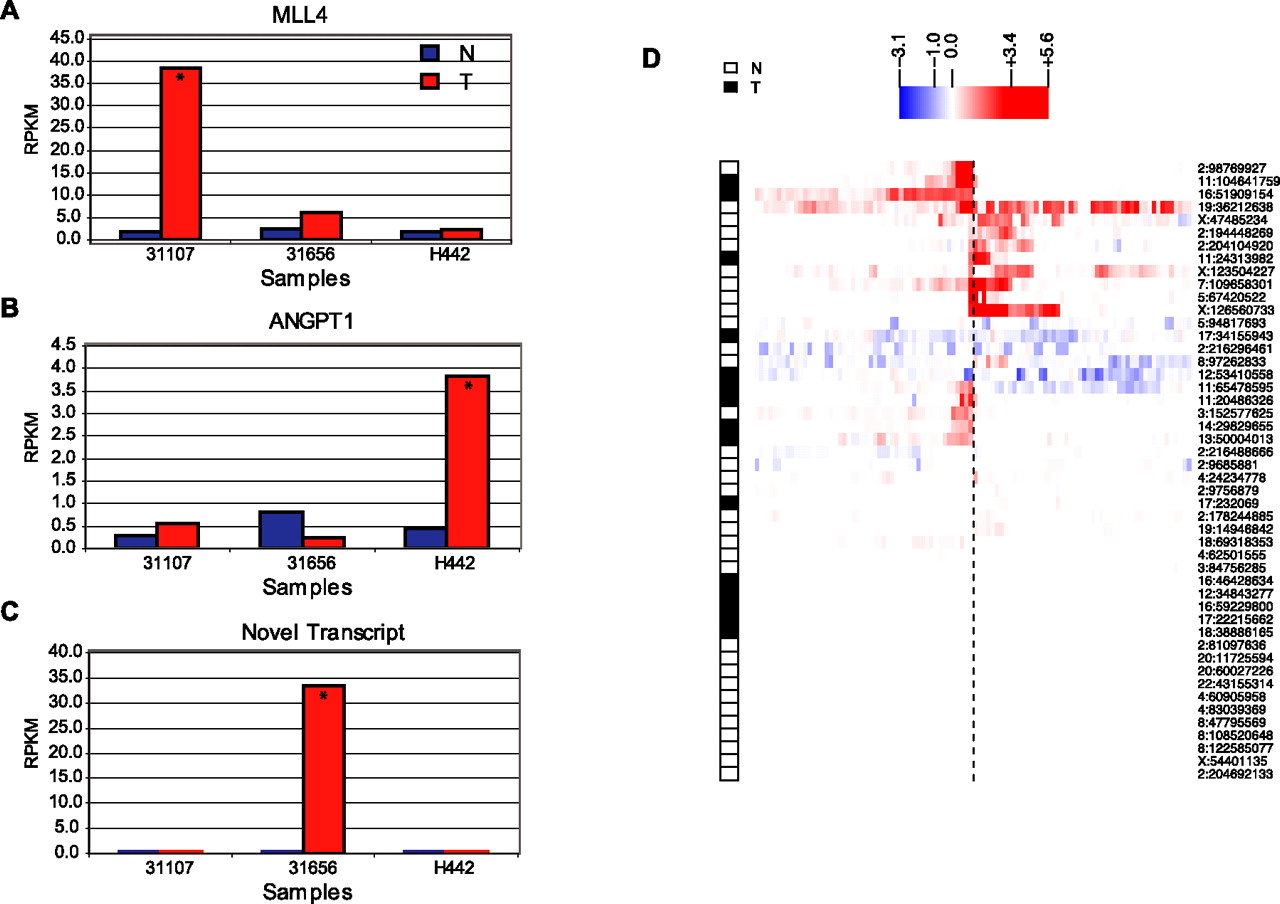
Transcriptional effect of HBV integration on the human genome. Local transcriptional effect of HBV integration on the human genome is shown, for the most abundant integration site for each patient (A–C) and for all integration sites (D), based on RNA-seq data. (A) MLL4 is highly overexpressed in the tumor sample with the HBV integration event (31107). (B) Substantial overexpression of ANGPT1 in the tumor from patient H442 with HBV integration upstream (∼10 kb) of ANGPT1. (C) A novel human-viral fusion transcript in the tumor sample with HBV integration in a nongenic region (patient: 31656). Asterisks indicate samples with viral integration. (D) Transcriptional effect at human-viral junctions defined by DNA-seq. The human-viral junctions supported by multiple DNA-seq chimeric reads (n = 48) are represented as a dotted line at the center. We then used the RNA-seq data to infer the transcriptional changes on each side of these junctions. The color in the heatmap represents the fold-change for each interval, measured as the difference in the generalized log of the RPKM of the altered genome (i.e., the genome containing the viral insertion) versus the unaltered genome. Samples carrying the insertion are indicated as either N (Nontumor) or T (Tumor). The rows were grouped by hierarchical clustering.