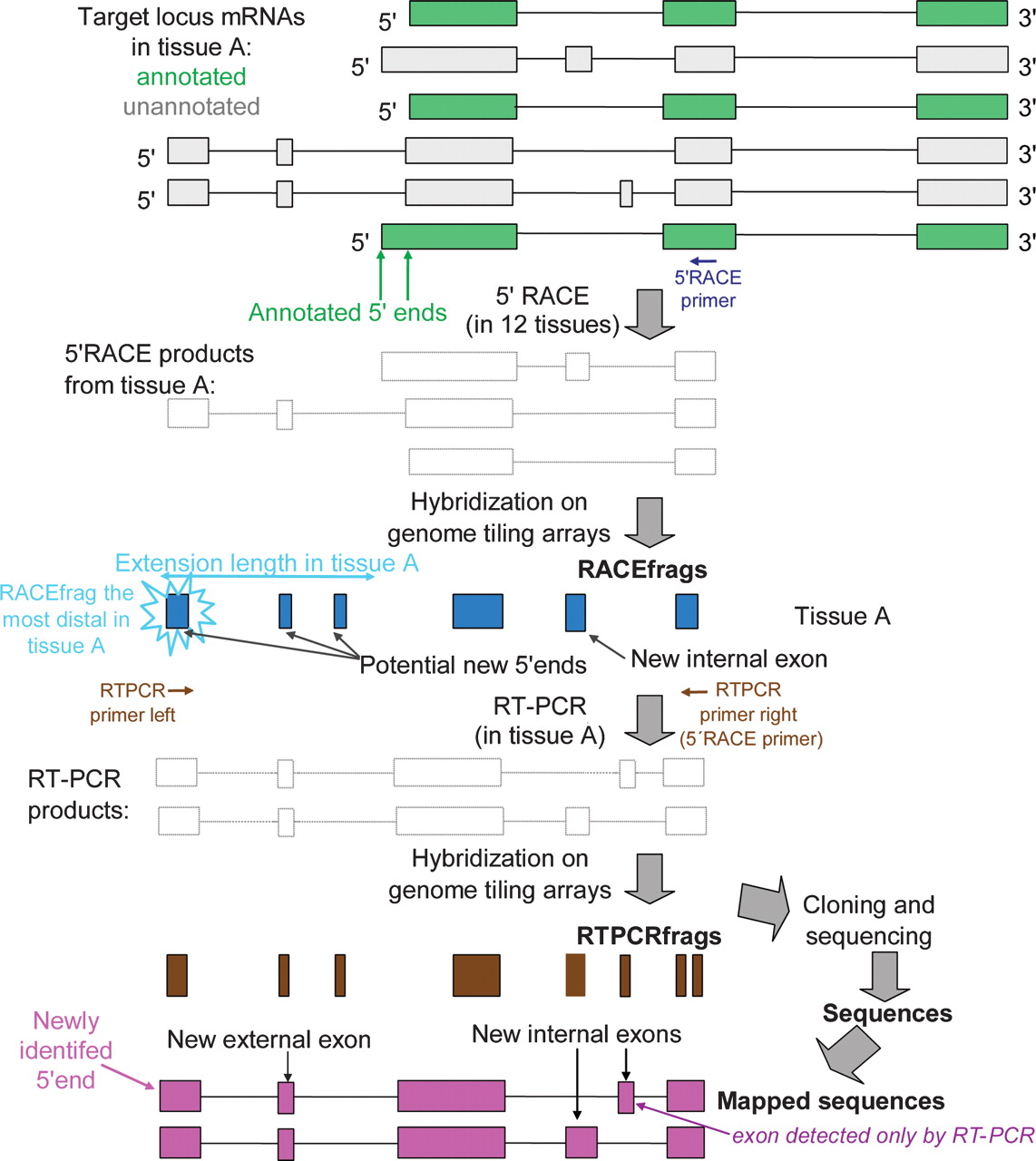
Schematic comparison of RACEfrags and RT-PCRfrags with annotated and unannotated transcripts. The locus to be interrogated is transcribed in alternatively spliced annotated (green) and unannotated (gray) isoforms. Rapid amplification of 5′ cDNA ends (5′ RACE) with a primer (blue arrow) mapping to a coding exon common to most of the transcripts (the index exon) results in a mix of cDNAs (ghost transcripts), which are hybridized to high-resolution tiling arrays to detect “RACEfrags” (blue boxes). RACEfrags are transcribed fragments specifically linked to the targeted coding locus. The connectivity between a RACEfrag overlapping an unannotated exon and the index exon can be verified by RT-PCR with two specific primers (brown arrows). This reaction produces a combination of overlapping alternatively spliced transcripts (ghost transcripts) that identify “RT-PCRfrags” upon hybridization to the same tiling array (brown boxes). Thus, RT-PCRfrags are transcribed fragments that link two targeted exons. Alternatively, these transcripts can be cloned and sequenced to precisely determine the beginning and the end of the novel exons and the exon composition of the transcripts (purple boxes). Because tiling arrays interrogate only nonrepeated regions and as they have a 20-bp resolution, RACEfrags and RT-PCRfrags do not fully overlap exons.