Picture this—you have just mapped a human disease locus to a particular region of a chromosome. With a click of a computer button, the region of chromosomal synteny in the zebrafish (Danio rerio) genome is revealed. Behold, there are several mutant zebrafish loci mapped in this general region of synteny. Another click and you find a fish mutant resembling your human disease. Further clicking reveals several independent alleles with varying phenotypes establishing the pathophysiology of the human disease. Does this sound farfetched? Well, recently several zebrafish mutants with “human” diseases have been found. With more infrastructure for the zebrafish system, the above scenario could become commonplace.
The zebrafish is an excellent system for developmental biologists and geneticists (Westerfield 1989; Detrich et al. 1999). The externally developing embryos are clear, allowing visualization of organ systems. The 1-inch size of the zebrafish allows large numbers of these vertebrates to be maintained in a relatively small space. In addition, each female lays >200 eggs per week. This enables the study of large numbers of meioses for positional cloning purposes. The genetic map has been continually improving over the past 2 years, and currently >2000 microsatellite markers and up to 400 genes have been defined (Knapik et al. 1998; Postlethwait et al. 1998) for the 1.7 × 109-bp genome (M. Fishman and J. Postlethwait, unpubl).
The zebrafish system was originally envisioned to provide important clues to normal embryogenesis and organ development. Because it is a vertebrate, the organism would bridge the gap betweenDrosophila/Caenhorhabditis elegans and mouse/human genetics. A flurry of candidate gene-cloning experiments revealed that the organism is an attractive one for developmental biology and clearly demonstrates the use of zebrafish for establishing embryonic axis and early neurogenesis (Solnica-Krezel 1999). Another hope for the system was that the vertebrate zebrafish would relate to the human, and mutants could define disease loci.
Positional cloning approaches in the zebrafish have been made possible by the development of key reagents such as YAC, PAC, and BAC libraries (Amemiya et al. 1999), as well as radiation hybrid panels (Kwok et al. 1998; M. Ekker, unpubl.). The first positional cloning project involved the isolation of the one-eyed-pinhead gene (Zhang et al. 1998), a novel cell surface molecule with EGF repeats. The second positional cloning project involved the isolation of the genesauternes (sau) (Brownlie et al. 1998). Saumutants have a normal number of blood cells circulating on day 2, but these blood cells fail to make hemoglobin. This mutant phenotype proved to be due to a defect in the erythroid synthase δ-aminolevulinate synthase (ALAS-2) gene, which regulates the first step in heme biosynthesis in embryonic red cells. Human patients withALAS-2 mutations have a disease very similar to the fish called congenital sideroblastic anemia, establishing this zebrafish mutant as the first animal model of this human disease (see Fig.1). Additionally, Shuo Lin and coworkers have provided evidence that the yquem (yqe) mutant is due to uroporphyrinogen decarboxylase (UROD) deficiency (Wang et al. 1998). This fish has the equivalent of human porphyria and further establishes the case that some of the zebrafish mutants will represent human diseases.
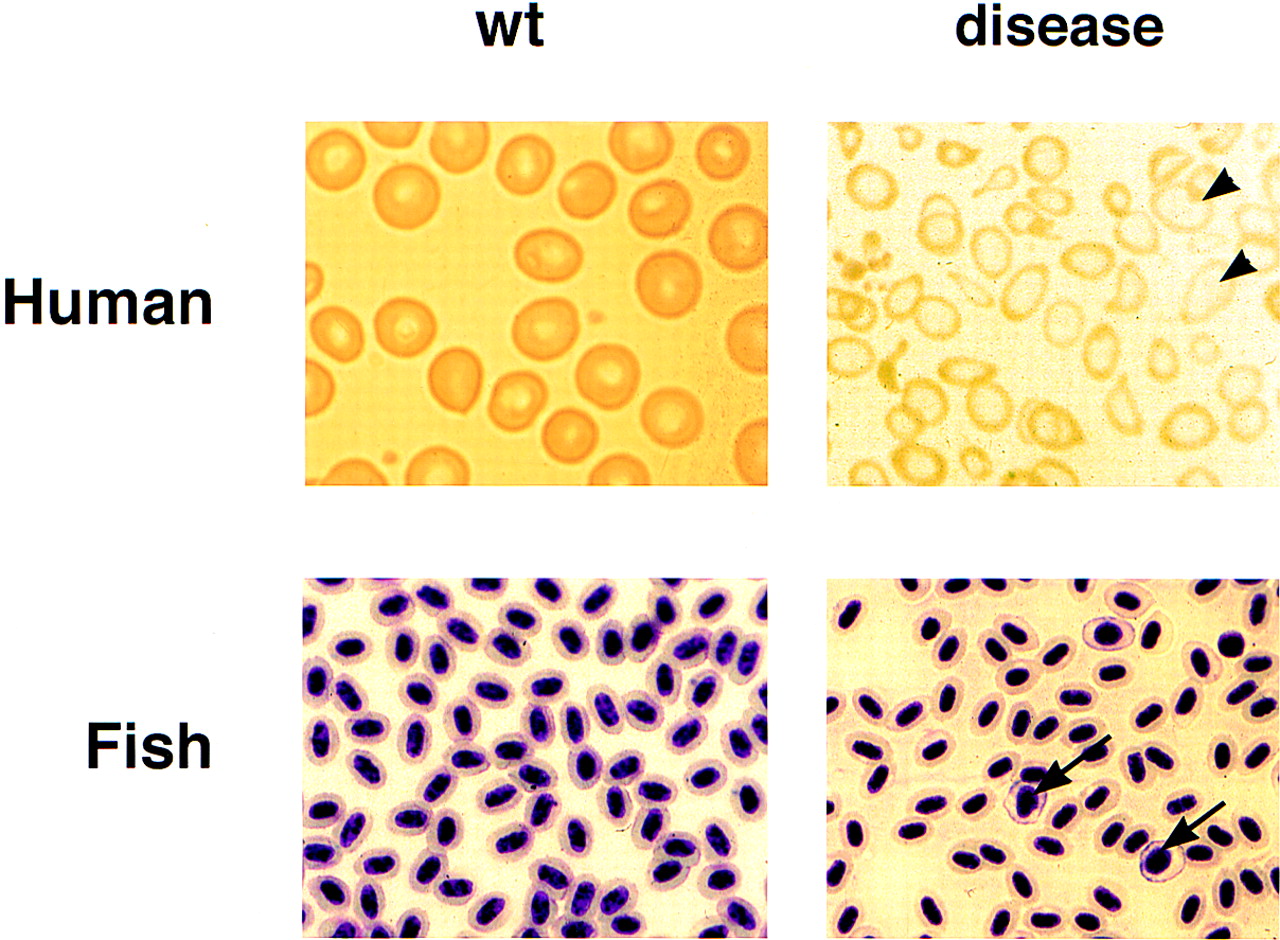
Comparison of the human and zebrafish diseases associated with mutations in ALAS-2. The red blood cells in the peripheral blood are normally round and uniform. In human congenital sideroblastic anemia, cells have abnormal shapes and are hypochromic (due to a low level of hemoglobin). Red cell precursors, such as reticulocytes, are larger (arrowheads). The increase in reticulocytes indicates a response of the marrow to the anemia. A similar hypochromia and increase in precursors is evident in the peripheral blood smear of sauhomozygous mutants (arrows).
Are the blood mutants unique among the zebrafish as to their relevance to human disease? Clearly, there are other phenotypes among all the zebrafish mutants that resemble human disorders (Driever and Fishman 1996). For instance, the zebrafish gridlock mutant has a defect similar to coarctation of the aorta in humans (Weinstein et al. 1995). In addition, there are zebrafish mutants with cystic kidneys that may represent polycystic kidney disease of humans (Drummond et al. 1998). It remains for clinicians to examine the zebrafish issue of Development (1996) to see whether other phenotypes resemble interesting diseases.
It was known previously that the mouse and human genomes share large blocks of chromosomal synteny, but no one believed that the fish chromosomal structure would resemble that of the human. For many chromosomal loci, the synteny is obvious between the fish and the human (Postlethwait et al. 1998). This facilitates positional cloning of the zebrafish genes, which can utilize information from the Human Genome Project. A zebrafish researcher can scour the human databases and look for candidate genes in the region near a zebrafish mutation. In the future, it should be possible for investigators studying human genetics to be able to interface directly to a zebrafish Web site (The Zebrafish Server, The Fish Net, ZFIN, http://zfish.uoregon.edu/) and evaluate mutants in a region of interest to the investigator. This process of “genome ping-ponging” based on these syntenic relationships will further establish the usefulness of the zebrafish for understanding human disease.
The article by Davidson et al. in this issue demonstrates the power of zebrafish to examine conserved genes and genome structure among the vertebrates. The GDF genes encode critical growth factors that signal developmental processes such as cell fate determination (Lee et al. 1998) and organ development (McPherron et al. 1997; Wolfman et al. 1997). By placing candidate genes on the genetic map, it should be possible to look for colocalization with mutant map positions. In this regard, large-scale mapping efforts have begun in the zebrafish. Two independent radiation hybrid panels have been created (Kwok et al. 1998; M. Ekker, unpubl.) and the mapping of >5,000–10,000 zebrafish genes is under way. This will provide excellent resource for developmental biologists, geneticists, as well as disease specialists. The future appears bright for the zebrafish.
Notes
[1] E-MAIL [email protected]; FAX (617) 355-7262.
REFERENCES
- ↵C.T. AmemiyaT.P. ZhongG.A. SilvermanM.C. FishmanL.I. Zon(1999) Methods Cell Biol. 60:235–258.
- ↵A. BrownlieA. DonovanS.J. PrattB.H. PawA.C. OatesC. BrugnaraH.E. WitkowskaS. SassaL.I. Zon(1998) Nat. Genet. 20:244–250.
- ↵H.W.I. DetrichM. WesterfieldL.I. Zon(1999) The zebrafish: Genetics and genomics. (Academic Press, San Diego, CA).
- ↵Development. 1996. 123..
- ↵W. DrieverM. Fishman(1996) J. Clin. Invest. 97:1788–1794.
- ↵I. DrummondA. MajumdarH. HentschelM. ElgerL. Solnica-KrezelA. SchierS. NeuhaussD. StempleF. ZwartkruisZ. RanginiW. DrieverM. Fishman(1998) Development 125:4655–4667.
- ↵E. KnapikA. GoodmanM. EkkerM. ChevretteJ. DelgadoS. NeuhaussN. ShimodaW. DrieverM. FishmanH. Jacob(1998) Nat. Genet. 18:338–343.
- ↵C. KwokR.M. KornM.E. DavisD.W. BurtR. CritcherB.H. PawL.I. ZonP.N. GoodfellowK. Schmitt(1998) Nucleic Acids Res. 26:3562–3566.
- ↵K. LeeM. MendelsohnT. Jessell(1998) Genes & Dev. 12:3394–3407.
- ↵A. McPherronA. LawlerS. Lee(1997) Nature 387:83–90.
- ↵J. PostlethwaitY. YanM. GatesS. HorneA. AmoresA. BrownlieA. DonovanE. EganA. ForceZ. Gong(1998) Nat. Genet. 18:345–349.
- ↵L. Solnica-Krezel(1999) Curr. Top. Dev. Biol. 41:1–35.
- ↵H. WangQ. LongS.D. MartyS. SassaS. Lin(1998) Nat. Genet. 20:239–243.
- ↵B. WeinsteinD. StempleW. DrieverM. Fishman(1995) Nat. Med. 1:1143–1147.
- ↵M. Westerfield(1989) The zebrafish book; A guide for the laboratory use of zebrafish ( Brachydanio rerio ). (University of Oregon Press, Eugene, OR).
- ↵N. WolfmanG. HattersleyK. CoxA. CelesteR. NelsonN. YamajiJ. DubeE. DiBlasio-SmithJ. NoveJ. SongJ. WozneyV. Rosen(1997) J. Clin. Invest. 100:321–330.
- ↵J. ZhangW. TalbotA. Schier(1998) Cell 92:241–251.