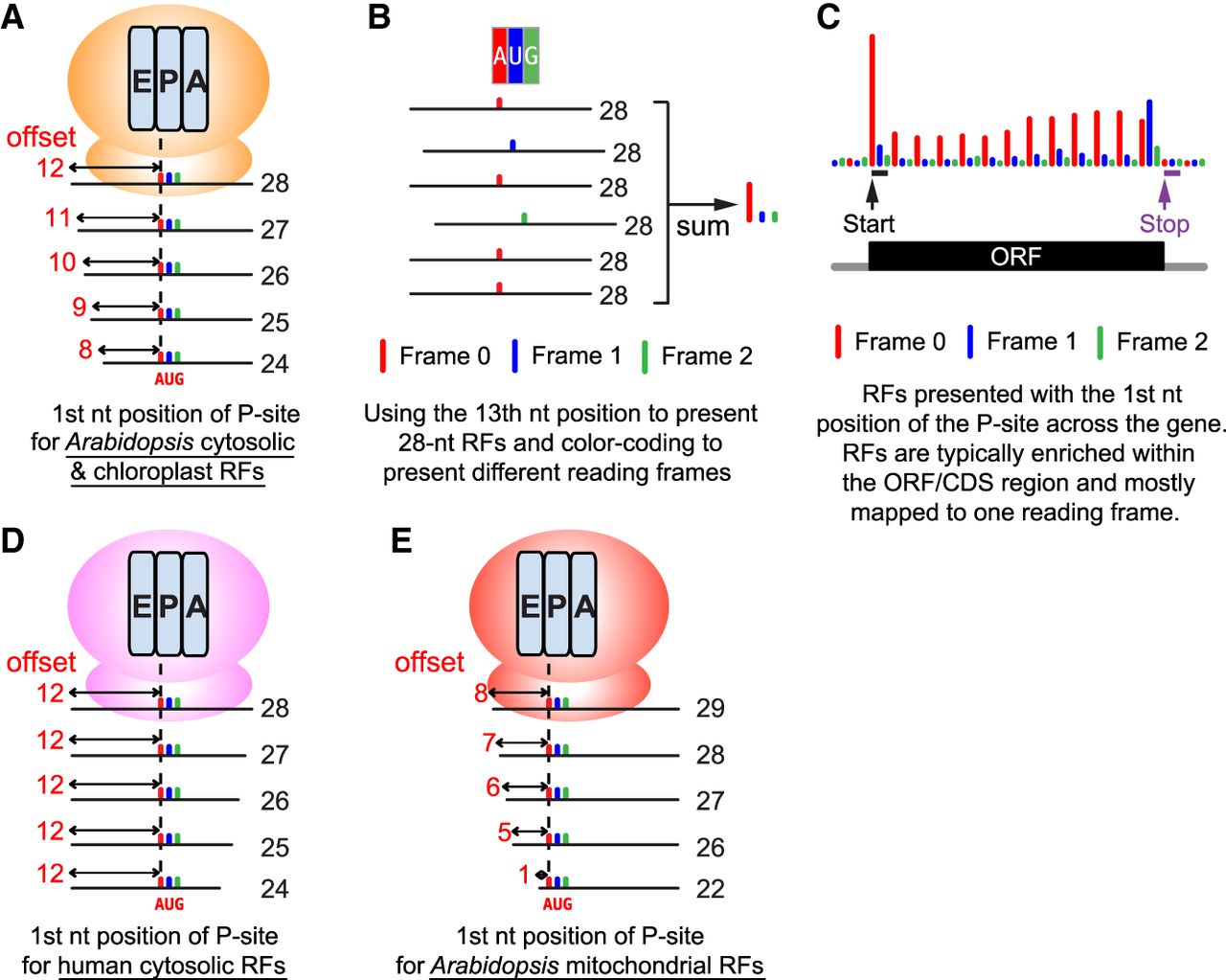
An illustration of how ribosome footprints (RFs) can be represented by a single nucleotide (nt) and color-coded to highlight 3 nt periodicity. (A) Metagene analysis by aligning all RFs mapped to the annotated start codon (AUG) can reveal the position of the P-site within ribosomes for each length of RFs. For Arabidopsis cytosolic and chloroplast RFs, the 13th nucleotide (offset by 12 nt from 5′) is typically the first nucleotide position of the P-site for a 28 nt RF. Similarly, the 12th nucleotide (offset by 11 nt from 5′) is the first nucleotide position of the P-site for a 27 nt RF. Note the P-site offsets could vary in different organisms and organelles (Ingolia et al. 2011; Hsu et al. 2016; Wu et al. 2024a). For high-quality Arabidopsis Ribo-seq data, ≤28 nt footprints exhibit excellent frame enrichment, making their P-site assignments reliable. In contrast, ≥29 nt footprints exhibit poorer frame enrichment (Wu et al. 2024a). Accordingly, for these Arabidopsis data, only ≤28 nt footprints are used for visualization, although longer footprints are still useful for other analyses. (B) An illustration showing how 28 nt RFs are mapped to a codon. The 28 nt RFs are presented by their 13th nucleotide position and color-coded by reading frame (annotated frame = frame 0, red; frame 1, blue; frame 2, green). Summing the reads mapped to each of the three nucleotides reveals the enrichment in frame 0 (red), which is expected in high-quality data sets. (C) Combining all RFs mapped to a particular gene produces a P-site plot, which visualizes the distribution of the RFs and colored 3 nt periodicity across the gene. At the codon prior to the stop codon, ribosome conformational change at the translation termination causes RFs to have a different length and results in an enrichment of frame 1 (blue) (Wu et al. 2024a). (D) The P-site offsets for human “cytosolic” RFs from induced pluripotent stem cells (iPSCs) and cardiomyocytes (Chen et al. 2020). Note that RNase digestion for different lengths of RFs occurs at the 3′ of the RFs in humans compared with the 5′ of the RFs in Arabidopsis (see A). (E) The P-site offsets for Arabidopsis “mitochondrial” RFs. The P-site offsets are shorter than the cytosolic and chloroplast RFs in Arabidopsis (see A).