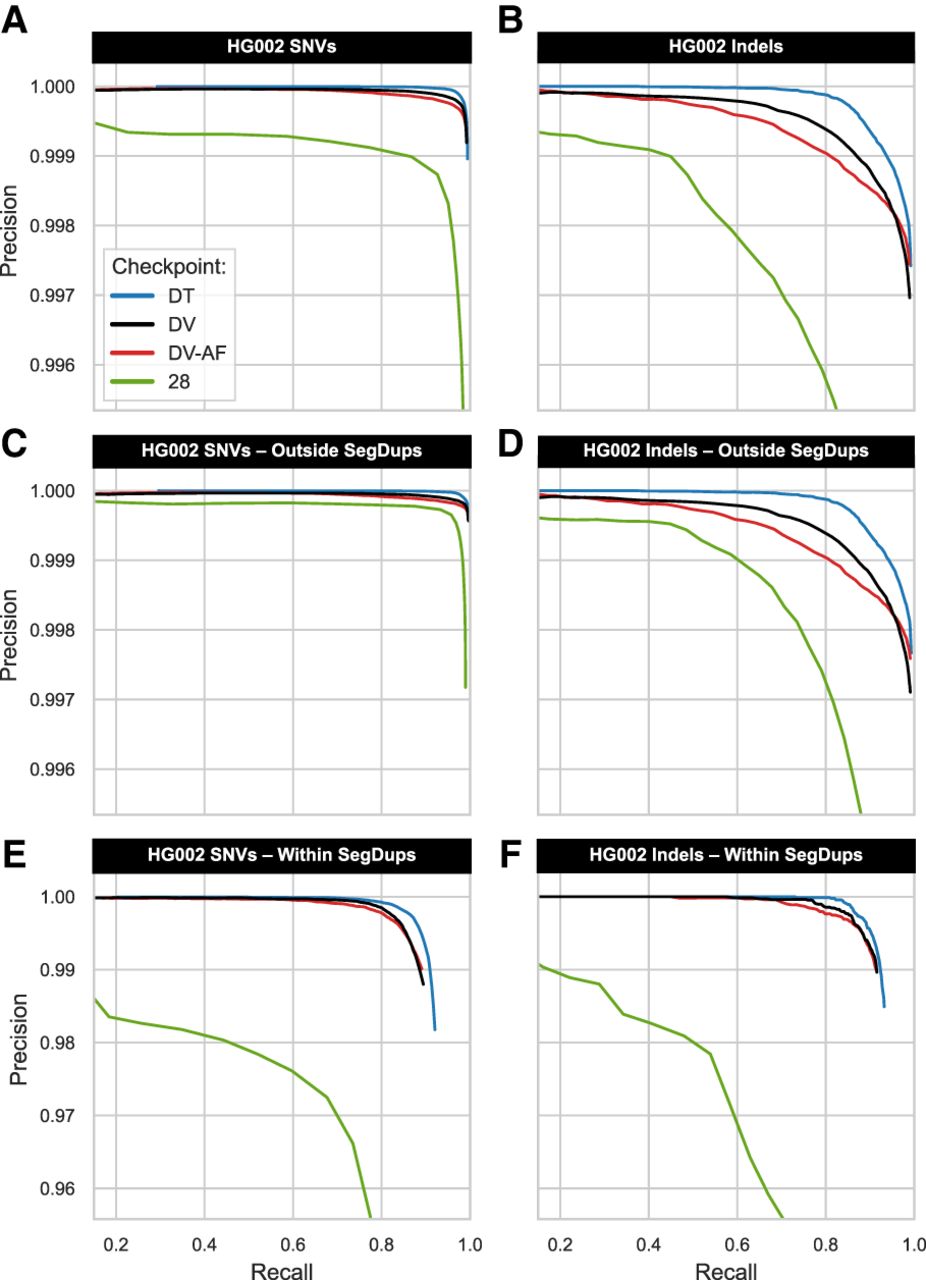
Benchmarking checkpoints with HG002. We used hap.py to calculate the precision-recall curve in a single human genome (HG002), where values above 0.5 indicate skilled prediction. We contrast three human-trained versions of DeepVariant (DV, DV-AF, DT) against a bovine-trained DV-AF checkpoint (28) that generalized well in humans and cattle. The top panels stratify genome-wide classification accuracy in SNV (A) or indels (B), controlling for the variable number of genotypes to allow direct model comparison. The middle panels use the GIAB stratifications (v3.5) to compare classification accuracy outside known segmental duplication (SegDup) regions in SNV (C) and indels (D). Note that the y-axis has a different scale for the bottom panels for stratification within SegDups in SNV (E) and indels (F). Models exclusively trained with human genomes outperform the bovine-trained model in HG002. However, the TrioTrain checkpoint's lower performance in repetitive regions is expected because bovine SegDups are not characterized to the same extent as in humans. Outside of known SegDups in HG002, the bovine-trained checkpoint created with TrioTrain is nearly perfect in SNVs (C). We infer that training with bovine genomes alters DeepVariant's priors for heterozygosity and copy number variation; these adjustments contribute to the marginally lower curve observed in human genome-wide precision and recall (top).