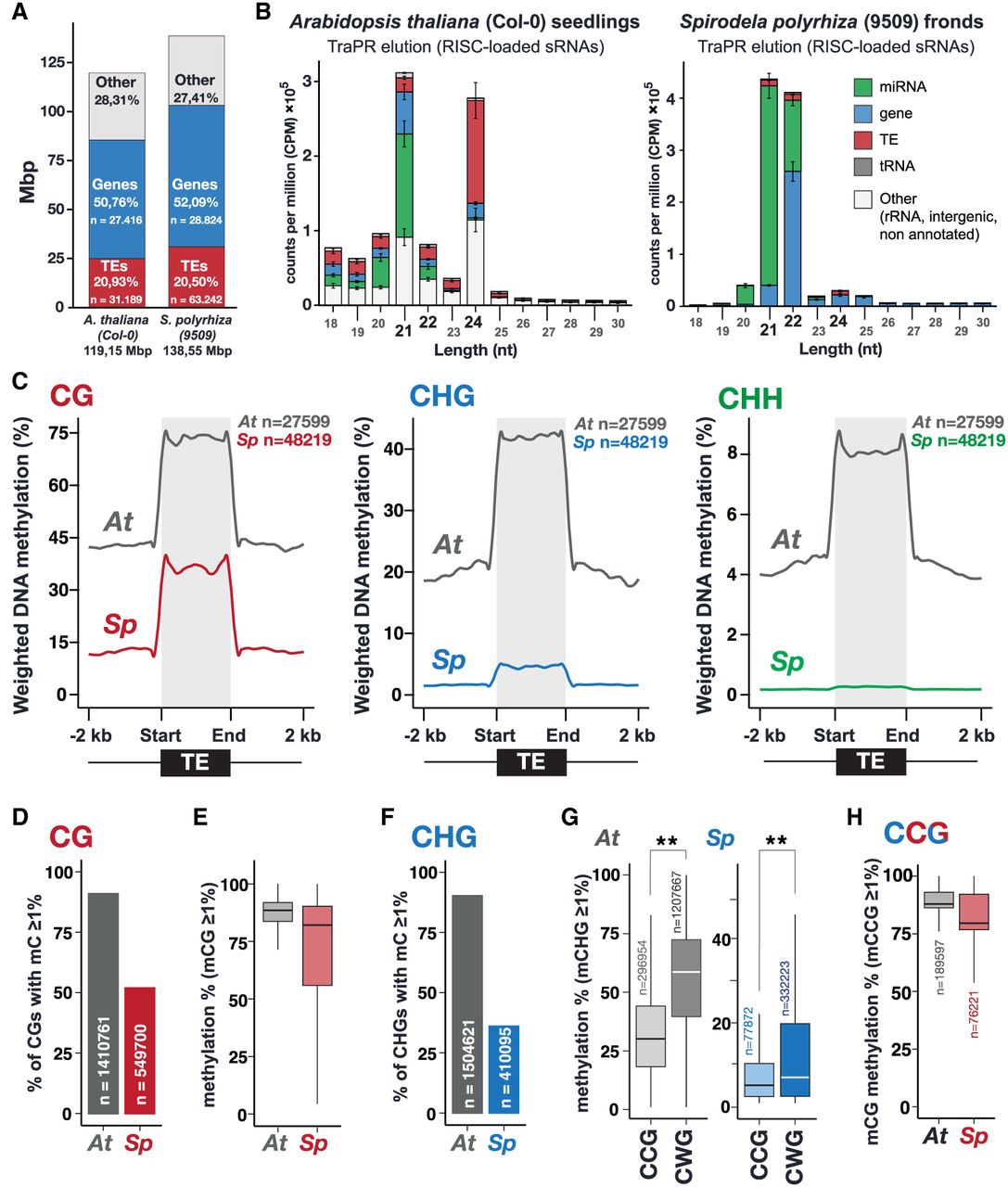
Spirodela polyrhiza sRNA and DNA methylation patterns. (A) Genome size, number, and genomic occupancy of genes and TEs in Spirodela and Arabidopsis. (B) Size distribution and genomic feature annotation of TraPR-purified sRNA from Arabidopsis and Spirodela. Error bars represent SD of the mean of three biological replicates. (C) Metaplots of averaged weighted DNA methylation in all three contexts (CG, CHG, and CHH, where H is any but G) over transposons (TEs; ≥100 bp) and 2 kb flanking regions in Arabidopsis and Spirodela. (D) Number and percentage of transposon CG sites displaying methylation levels ≥1% in Arabidopsis and Spirodela. (E) DNA methylation levels at CG sites from D. (F) Number and percentage of transposon CHG sites displaying methylation levels ≥1% in Arabidopsis and Spirodela. (G) DNA methylation levels at CCG and CWG in Arabidopsis and Spirodela. (H) Inner 5mC (CmCG) DNA methylation levels at CCG sites with outer 5mC (mCCG) ≥1% in Arabidopsis and Spirodela. (**) P-value < 2.22 × 10−16 (Wilcoxon rank-sum test). In all box plots, the median is indicated by a solid bar; the boxes extend from the first to the third quartile; and whiskers stretch to the furthest value within 1.5 times the interquartile range. (At) Arabidopsis, (Sp) Spirodela.