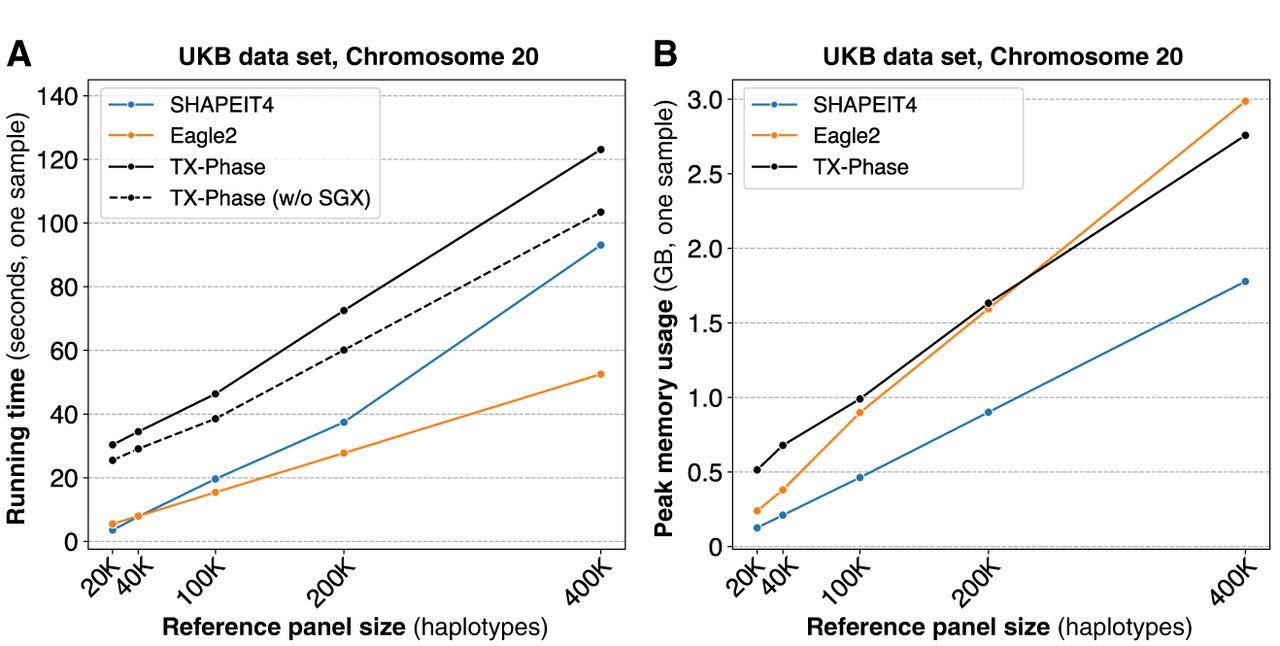
TX-Phase maintains practical running times for phasing individual samples while providing enhanced security. We report the per-sample running times (A) and peak memory usages (B) of TX-Phase, SHAPEIT4, and Eagle2 for phasing the UKB genotyping array data set (Chromosome 20; 19,959 variants) with varying reference panel sizes up to 400,000 haplotypes. Additionally, we plot the running times of TX-Phase without SGX for comparison, in which our program is executed in a conventional (unprotected) computing environment. The results demonstrate that TX-Phase effectively minimizes the overhead of side-channel resilient computation in TEEs, achieving practical performance comparable to existing tools, or is only marginally more costly (e.g., by less than a factor of three) on large reference panels.