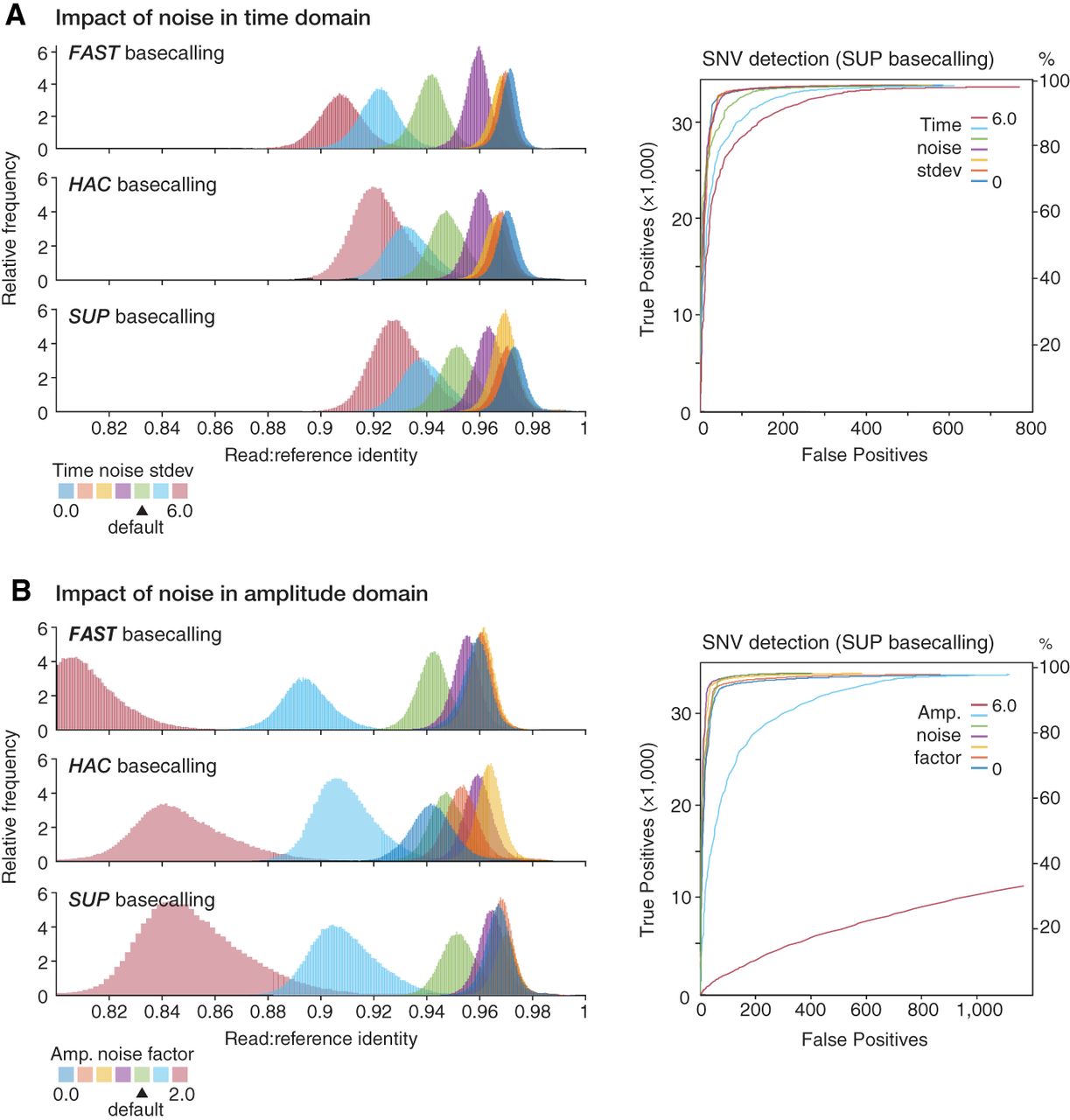
Modeling the impact of time and amplitude noise on sequencing accuracy. (A, left) Guppy basecalling accuracy, as measured by read:reference identity score distributions, for repeated experiments in which warping in the time domain (i.e., variation in the DNA translocation speed) was varied, whereas other parameters were held at default. Higher values indicate increasing noise and default value is ‐‐dwell-std = 4 (green distribution). Experiment was repeated with Guppy's FAST (upper), HAC (middle), and SUP (lower) basecalling models. (Right) ROC curves assess accuracy of SNV detection by Clair3 on the same data sets (colors are matched). (B) Same analysis as above but with noise in the amplitude domain (‐‐amp-noise) varied, whereas other parameters are static.