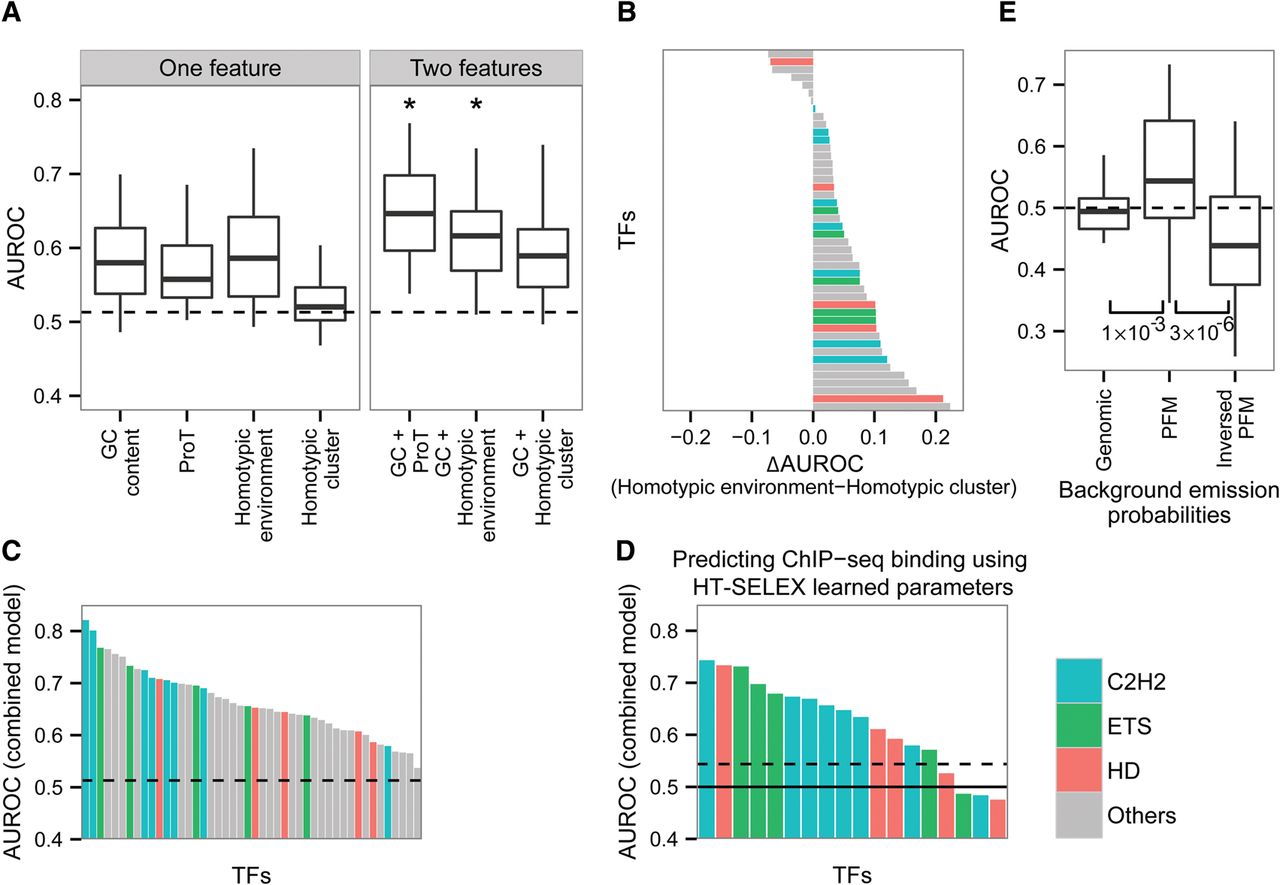
Predicting bound and unbound TF motifs. (A) L2-regularized multiple linear regression models based on one or two features in vivo. The features characterizing the average GC content, the average propeller twist (ProT), the average PFM similarity scores (homotypic environment), and the sum of all significant PFM similarity scores (using FIMO P-value cutoff of 0.001; homotypic cluster). All features were extracted from 300 bp upstream of and downstream from the core motif, excluding the core motif. Box plots represent the distribution of the AUROC for all TFs using one or two features. The dashed line represents the maximum AUROC obtained using randomly shuffled data. Asterisks are shown for features in which the AUROC obtained using the two-feature model is significantly higher than the AUROC obtained using each feature separately. (B) For each TF, comparison of the AUROC obtained using the homotypic environment model and the homotypic cluster model. The TFs are colored according to the color code used for TF families: cyan for C2H2 TFs, green for ETS TFs, red for homedomains, and all others in gray. (C) AUROC values for each of the TFs, employing a model that incorporates the best preforming features: GC content, propeller twist, and homotypic environment. Dashed line represents the maximum AUROC obtained using randomly shuffled data. (D) AUROC of the combined model that was trained using the in vitro data and was tested on the in vivo data. Dashed line represents the maximum AUROC obtained using randomly shuffled data. Solid line shows AUROC of 0.5. (E) AUROC of the HMMs using different emission probabilities for the background state: the genomic nucleotide frequency, average nucleotide frequency of the PFM, and the inversed average nucleotide frequency of the PFM. Wilcoxon test P-values are shown below. The dashed line represents AUROC of 0.5.