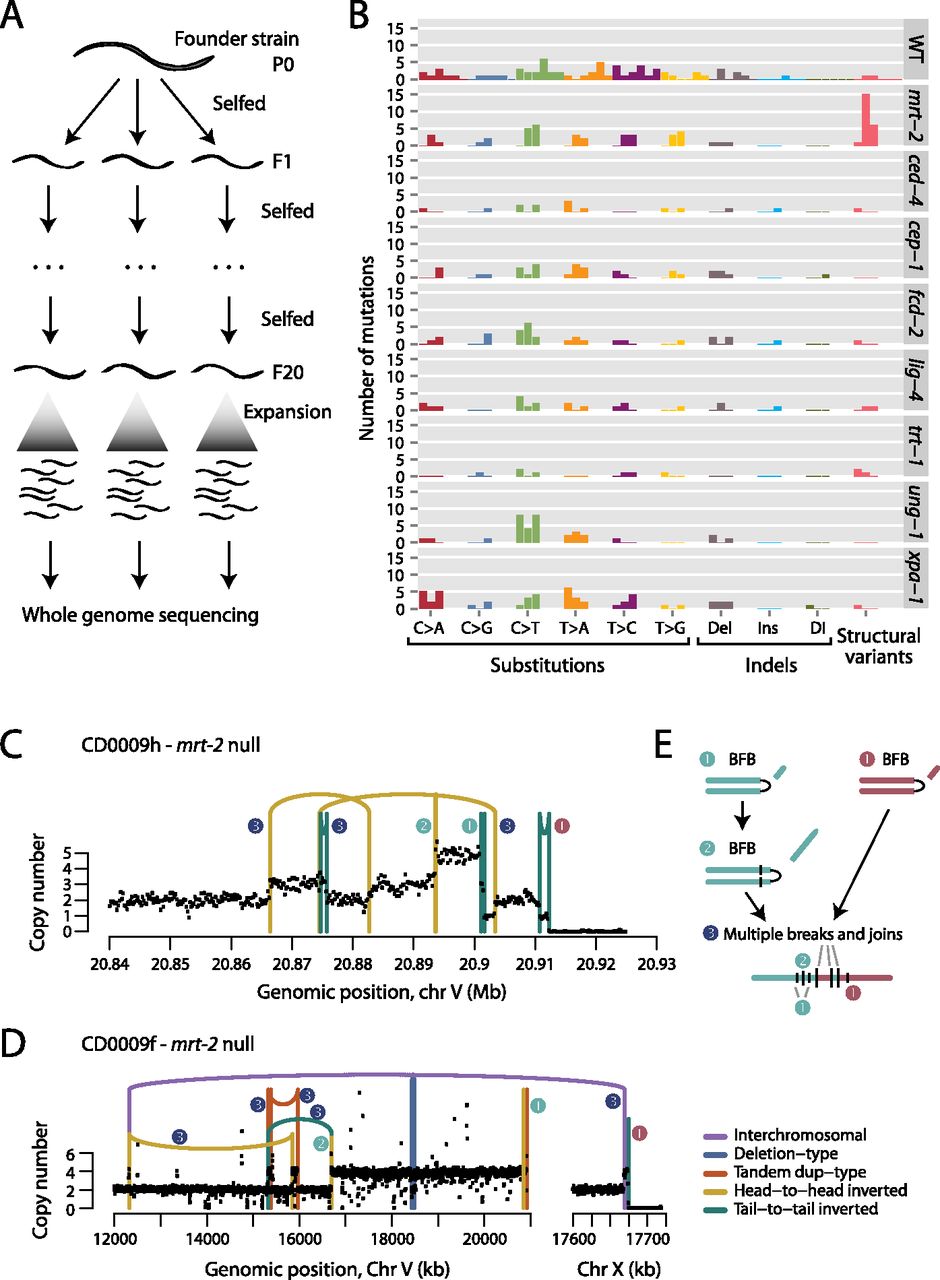
Mutations in C. elegans followed for 20 generations. (A) The experimental design used three progeny of a founder or parental (P0) animal, propagated by self-fertilization for 20 generations (F1: filial 1; F20: filial 20). The 20th generation worm was expanded to generate sufficient DNA for whole-genome sequencing. (B) Numbers and distribution of acquired mutations across wild-type (WT) and different genetic knockout lines. Each replicate worm is shown as a separate bar, colored by the class of variant. (Del) Deletion; (Ins) insertion; (DI) complex indel. (C) Copy-number profile and genomic rearrangements observed on chromosome V for an mrt-2 deficient animal. Numbers next to the genomic rearrangements refer to the inferred sequence of events. (D) Copy-number profile and genomic rearrangements observed on chromosomes V and X for another mrt-2-deficient animal. Numbers next to the genomic rearrangements refer to the inferred sequence of events. (E) Proposed model for the evolution of the copy-number changes and genomic rearrangements in C and D. A telomere crisis precipitates several breakage-fusion-bridge (BFB) cycles, which are resolved by a cell cycle in which multiple DNA breaks are simultaneously joined.