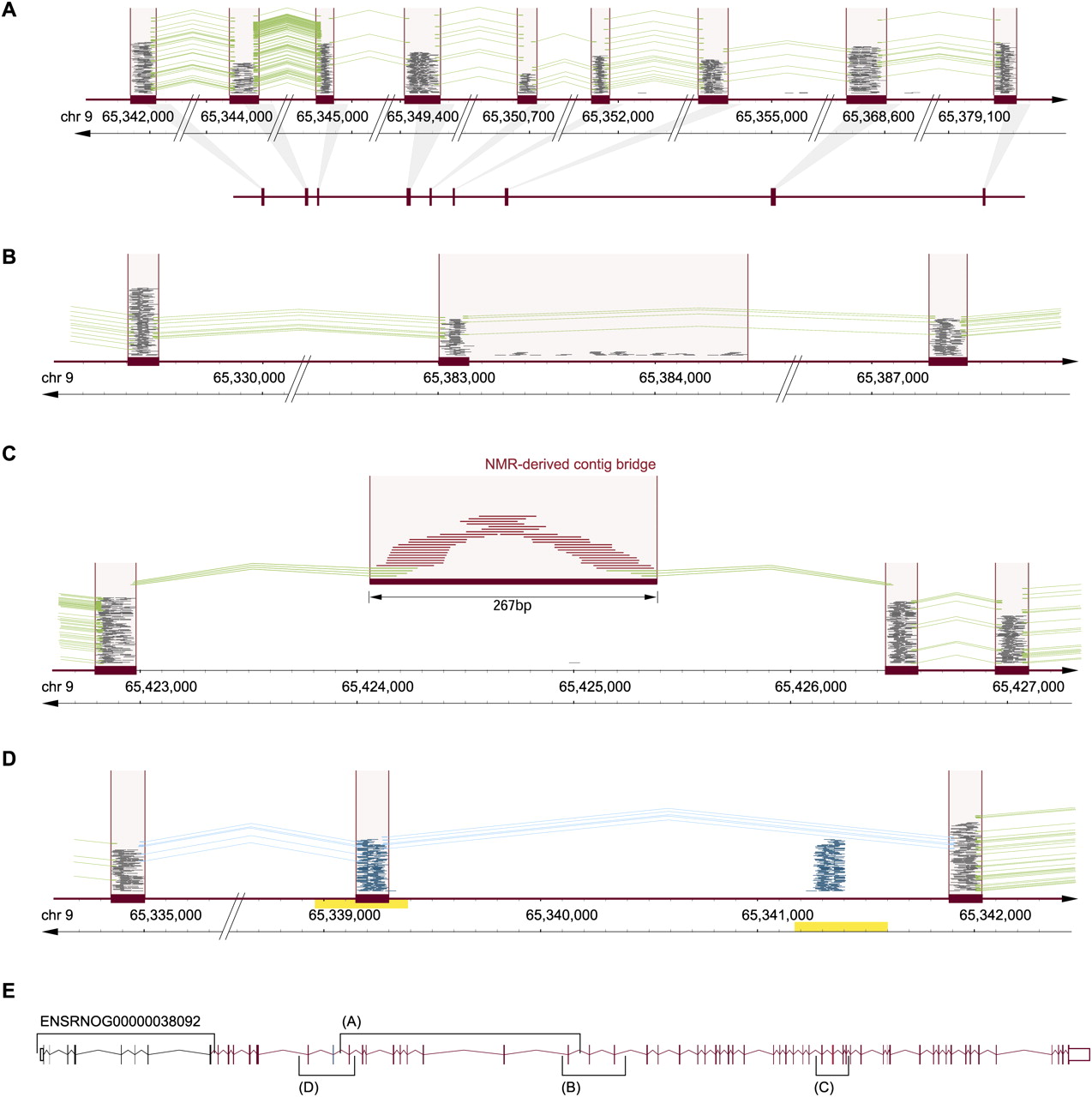
Integrating information from UMR, sjUMR, MMR, and NMR read classes for complex genome (re-)annotation. (A) UMR- and sjUMR-based definition of new exon groups. A novel gene candidate was identified on chromosome 9 through several UMR clusters (satisfying the classifier criterion of a read density > 4), which were connected through splice junctions defined by sjUMR. Expression of exons was coregulated (suppressed by 70% after SNL). An open reading frame encoding 409 amino acids was noted lacking a start and stop codon, suggesting that an incomplete gene fragment was identified consisting of nine exons. (B) sjUMR validation of a low-read density UMR cluster as a novel exon. Consistent with the limited sensitivity of the UMR cluster classifier (91%), a UMR read cluster with a read density of 3.9 was initially not classified as an exon (because its density was <4); but sjUMR-based SJs defined it unambiguously as a novel exon connecting the above gene fragment to a series of 18 3′-located UMR clusters. Note that the cluster density was low because a region of scattered intron reads was added to the read cluster by the sliding window, an imprecision that was rectified by agnostic splice site mapping, which defined precise exon borders. Through the step depicted in this panel, the candidate gene was extended to 28 exons. (C) NMR-derived contig bridge. A faulty or missing section in the reference genome can be indicated by runs of N (ambiguous bases) as encountered 3′ of the 28 exons assembled above (50 scattered N). Such missing sections in the reference are also an important reason why some mRNA-seq reads do not match. Accordingly, we found that in such cases, contigs of NMR can be assembled bridging faulty sequences, as shown here. As a result of the NMR contig-bridge, the gene candidate could be expanded to 49 exons. (D) MMR candidate exons discriminated by sjUMR. Exon copy numbers >1 in the reference sequence result in clusters of MMR (blue) matching collectively to more than one site. A case of two such MMR clusters in close proximity (within <2 kb; note duplicated region indicated by yellow bar) was observed 5′ of the above gene candidate. As shown here, the ambiguity was resolved by sjUMR; this led to a 5′ extension of the gene candidate by another 15 exons, which resulted in a complete gene with start and stop codon consisting of 64 exons. (E) A summary of the novel 64-exon gene is shown encoding a 3313-amino-acid-long protein (14,084-bp mRNA). While the study was under way, a homologous mouse protein with important CNS relevance was reported (“large previously unknown protein”) (Lu et al. 2009), termed UNC-80, which serves as a substance-P and neurotensin coreceptor. Down-regulation of the rat UNC-80 rat homolog after SNL may contribute to allodynia through alterations in peptide activity in the DRG.