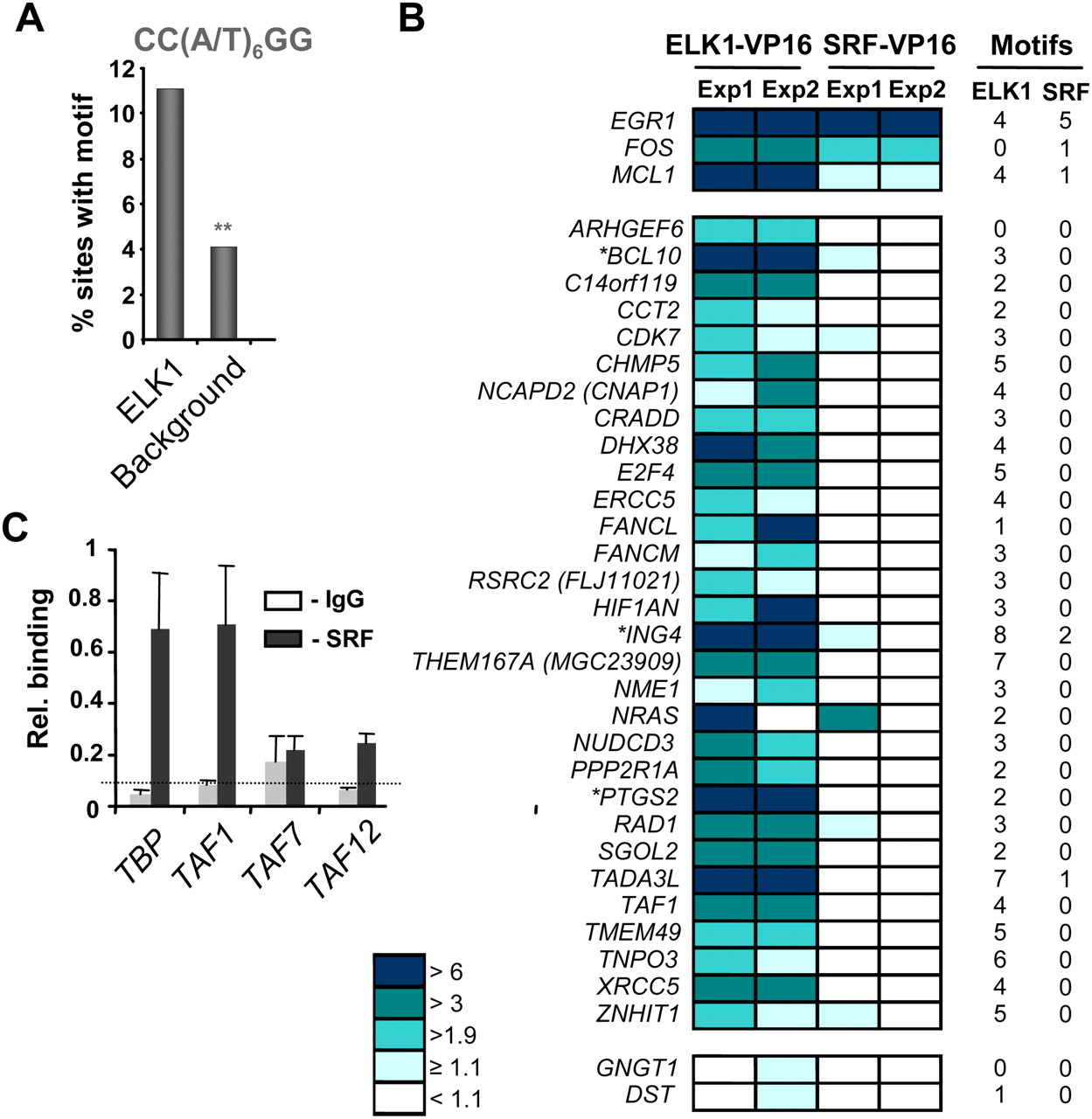
SRF-dependent and independent ELK1 target promoters. (A) Over-representation of the CC(W)6GG SRF-binding motif in the ELK1 FDR < 10 data set in comparison to a background data set. **P ≤ 1 × 10−4. (B) Reporter gene analysis in 293 cells of a panel of promoters containing ELK1-binding regions. Cells were cotransfected with the indicated reporter plasmids and plasmids encoding either ELK1–VP16 or SRF–VP16. Data are shown from two independent experiments (Exp1/2) and the levels of reporter activity are color coded according to fold activation by ELK1–VP16 or SRF–VP16. The numbers of ELK1 (nonredundant hexamers derived from CCGGAAGT motif) and SRF (CC(W)6GG) binding motifs in each promoter is shown on the right, and genes also found in the high confidence SRF ChIP-chip FDR < 1 data set are indicated by asterisks. (C) qPCR-ChIP analysis of SRF binding to the indicated promoters. The dotted line represents average binding across all IgG control precipitations. Data are the average of duplicate samples and representative of three independent experiments.