Compared codon usage in the whole genome and C-left vs. tRNA gene usage
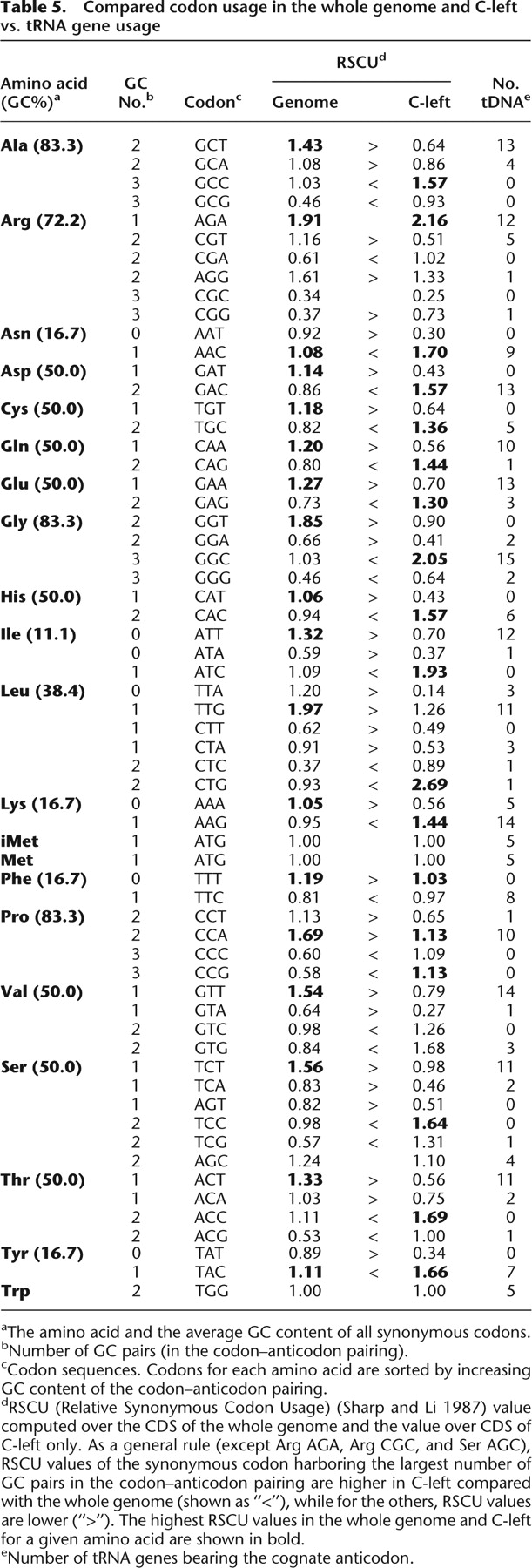
[i] aThe amino acid and the average GC content of all synonymous codons.
[ii] bNumber of GC pairs (in the codon–anticodon pairing).
[iii] cCodon sequences. Codons for each amino acid are sorted by increasing GC content of the codon–anticodon pairing.
[iv] dRSCU (Relative Synonymous Codon Usage) (Sharp and Li 1987) value computed over the CDS of the whole genome and the value over CDS of C-left only. As a general rule (except Arg AGA, Arg CGC, and Ser AGC), RSCU values of the synonymous codon harboring the largest number of GC pairs in the codon–anticodon pairing are higher in C-left compared with the whole genome (shown as “<”), while for the others, RSCU values are lower (“>”). The highest RSCU values in the whole genome and C-left for a given amino acid are shown in bold.
[v] eNumber of tRNA genes bearing the cognate anticodon.