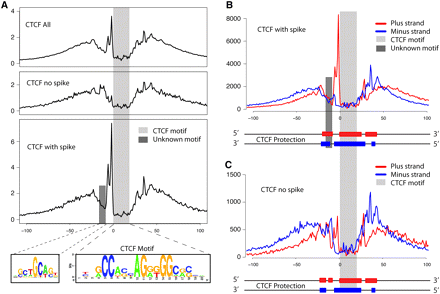
High-resolution analysis of CTCF binding sites. (A) Cumulative footprinting signal at all CTCF motif predicted sites that includes sites with and without a large increase in DNase I digestion upstream of the CTCF motif. The light gray bar indicates the location of the known CTCF motif. The dark gray bar represents the location of a novel binding motif. The novel binding motif was only detected in CTCF footprints that contain the small upstream region with a spike in DNase I hypersensitivity (HS). Note that the entire protected region is approximately 50–60 bases. (B) Strand-specific DNase-seq signal for the subset of CTCF motif identified sites that contain the upstream DNase I HS spike. The DNase I HS spike is only detected on the positive strand. (C) Similarly for the CTCF motif identified sites without the upstream DNase I HS spike. The diagram below each plot in B and C illustrates the estimated strand-specific protected regions surrounding the CTCF motif predicted sites.











